![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
71 Cards in this Set
- Front
- Back
|
What is the central dogma for Eukaryotes? |
(replication)DNA -->(transcription) pre-mRNA=> splicing=>mRNA(translation)--> protein |
|
|
typical eukaryotic gene image look at |
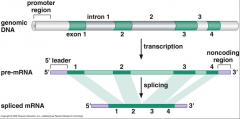
|
|
|
what is an exon? |
coding sequence in eukaryotic EX= exit the nucleus |
|
|
what is intron? |
the non-coding or intervening sequences in euk. genes IN = stay w/in the nucleus |
|
|
what is pre-mRNA? |
the primary transcription of DNA |
|
|
what is RNA splicing? |
the process of introns being removed from the pre-mRNA. |
|
|
What is spliceosome? |
a complex of specialized RNA & protein subunits that removes introns from pre-mRNA |
|
|
what is alternative splicing? |
process of the re-combination of different exons in eukaryotes. -Major source of genetic diversity of eukaryotes. |
|
|
Describe RNA splicing? |
-mRNA's contain different selection of eons can be generated from a given pre-mRNA. |
|
|
Name the 3 classes of RNA splicing pathways? |
1.) nuclear pre-mRNA splicing (spliceosome mediate intron removal) 2.) group 1 introns 3.) group 2 introns (intron itself folds into a specific conformation w/in the pre-mRNA & catalyzes the chemistry of released by itself and the exon ligation.)
|
|
|
How does the pre-mrNA splicing work? |
there are 2 steps involved: 1.) 1st step: find the splice sites 2.) 2nd step: intron is moved in a form called Lariat through 2 transesterification reactions as the flanking eons are joined. |
|
|
Name the splice sites in the first step of pre-mRNA splicing? |
1.) 5' splice site 2.) 3' splice site 3.) Branch point site |
|
|
what is 5' splice site? |
boundary at the 5' end of the intron is marked by a specific nucleotide sequences (GU-AG law) |
|
|
what is 3' splice site? |
boundary at the 3' end of the intron is marked by a sequence. |
|
|
what is brand point site? |
sequence w/in the intron & is followed by a polypyrimidine tract (Py tract) rich in pyrimidine. |
|
|
Sequences w/in the RNA deter time where what occurs? |
splicing |
|

Describe what is happening? |
-GU 5' splicing site, AG in 3' splicing site & A in branch point site are the most conserved sequences & they are all in the intron. -These sequences are important for the distinguish btw intron & exon, remove of intron, linkage of eons, & delineate where splicing will occur. |
|
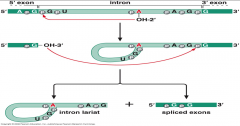
what step is this? |
second step of pre-mRNA splicing -Intron is removed through 2 trasesterificatiron reactions in a form called Lariat & the flanking eons are joined. |
|
|
what happens in the trasesterirication reaction 1? |
-part of second step -2'OH attacks 5' site -Result = the 5' exon is released -5' intron forms a 3 -way junction |
|
|
what happends in the transesterification reaction 2? |
- THE OH of the 5' exon attacks the phosphoryl grp at the 3' splice site. -RESUT: 5 ' and 3 ' exons are joined -intron is liberated as a lariat |
|
|
in the two transesterrification reactions, there is no what? |
net gain in the number of chemical bonds, so no energy is demanded by the process |
|
|
Why do we see a large amount of ATP being consumed during splicing reaction? |
this is energy is required to properly assemble and operate the splicing machinery, not for the chemistry. |
|
|
Splicesome is a complex of what? |
specialized RNA & protein subunits |
|
|
Splicesome comprises about how many proteins & RNA's? |
150 proteins; 5 RNAs |
|
|
Wha are small nuclear RNAs composed of? |
Five RNAs ( U1, U2, U4, U5, U6, 100-300 nt) |
|
|
The complexes of snRNA & proteins are called what? |
snRNPS (small nuclear ribonuclear proteins |
|
|
Spliceosome is the largest what? |
snRNP, and the exact make up differs at different statnges of the splicing reaction |
|
|
Spliceosome mediate what?s |
splicing of introns from pre-mRNA |
|
|
Many functions of the spliceosome are carried out by what? |
RNA components |
|
|
what are the 3 roles of snRNPS in splicing? |
1.) recognizing the 5' splice site & the branch site 2.) bringing those sites together 3.) catalyzing the RNA cleavage |
|
|
What are the 3 things that are important during splicing? |
1.) RNA-RNA 2.) RNA- PROTEIN 3.) protein-protein interactions |
|
|
what is Nuclear Pre-mRNA splicing? |
pathway medicated by splicesosome & involved in assembly, rearrangement & catalysis w/in the splicesome |
|
|
What happens in step 1 of nuclear pre-mRNA splicing pathway assembly? |
Step 1: U1 recognize 5 ' splice site -one subunit of U2AF binds to branch site, former subunits interact w/ BBP & helps bind to branch point. -Early (E) complex is formed |
|
|
What happens in step 2 of nuclear pre-mRNA splicing pathway assembly? |
Step 2: -U2 binds to branch site and then A complex = formed -Base-pairing btw the U2 & branch site is such that the branch site A is extruded. A residue is available to react w/ the 5 ' splice site. A COMPLEX |
|
|
What happens in step 3 of nuclear pre-mRNA splicing pathway assembly? |
Step 3: -U4, U5, and U6 form the trip-snRNP particle -w/ the entry of the tri-snRNP, the A complex is converted into b complex B COMPLEX |
|
|
What happens in step 4 of nuclear pre-mRNA splicing pathway assembly? |
Step 4: -U1 leaves the complex, U6 replaces it at the 5' splice site -U4 is released from the complex, allowing U6 to interact with -U2: this arrangement called "The C Complex" |
|
|
What happens in step 1 of nuclear pre-mRNA splicing pathway catalysis ? |
step 1: - C complex = active site, w/ U2 & U6 Rna's being brought together -allows 5' splice accomplish Transesterificaiton 1 |
|
|
What happens in step 2 of nuclear pre-mRNA splicing pathway catalysis ? |
Step 2: -U5 SnRNP helps to bring the two exon together -allows second transeterification reaction,
|
|
|
What happens in step 3 of nuclear pre-mRNA splicing pathway catalysis ? |
release of the mRNA product & snRNPS |
|
|
image of step 1 nuclear pre-mRNA splicing assembly |

|
|
|
image of step 1 nuclear pre-mRNA splicing assembly part two |
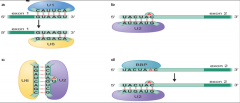
|
|
|
image of step 2 nuclear pre-mRNA splicing assembly |
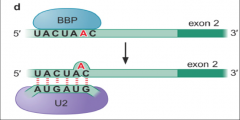
|
|
|
image of step 2 nuclear pre-mRNA splicing assembly part two |
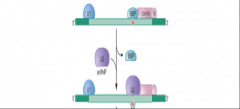
|
|
|
image of step 3 nuclear pre-mRNA splicing assembly |
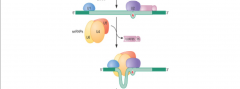
|
|
|
image of step 4 nuclear pre-mRNA splicing assembly |
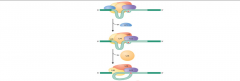
|
|
|
image of step 1 nuclear pre-mRNA splicing assembly |
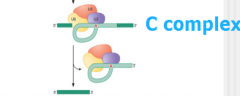
|
|
|
image of step 2 & 3 nuclear pre-mRNA splicing assembly |

|
|
|
image overview nuclear pre-mrNA splicing |
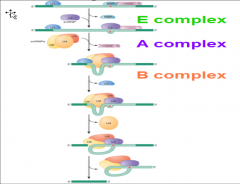
|
|
|
what is self-splicing? |
a few rare introns can remove themselves from pre-mRNA = self-splicing |
|
|
what are the two classes of self-splicing? |
Group I Group II |
|
|
what are the features of self-splicing? |
-does not need spliceosome machinery -Rna of intron mediates the chemistry of removal by intron itself folds into a specific conformation w/in the pre-mRNA and catalyzes the chemistry of released by itself & the exon ligation. (RNA enzyme endcoded by intron = riboenzyme)
|
|
|
what is group 1intron and group 2 intron in self-splicing? |
-Group 1 intron self-splicing releases a linear introns rather than a lariat -Group 2 intron self-splicing has the similar chemistry as the spliceosomemediated splicing |
|
|
overview of 3 RNA splicing pathways |
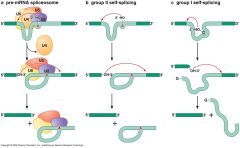
|
|
|
Features of alternative splicing? |
-process of re-combination of different exon in eukaryotes -many genes in higher eukaryotes encode RNA that can be spliced in alternative ways to generate 2 or more diff. mRNA's & diff. protein products (or isoforms) -Microarry analyses = 40% drosophila & 75% human genes = alternative splicing
|
|
|
Alternative splicing is a major what? |
source of genetic diversity in eukaryotes |
|
|
Name the ways of alternative splicing can occur? |
1.) Trans-splicing 2.) Two diff. mrNa's generate from a single gene = 2 diff. proteins 3.) mult. mRNA generated from a single gene = mult. proteins.
|
|
|
what is trans-splicing? |
2 exon from diff. RNA molecules can be joined together. -its generally rare, but it occurs in almost all of mRNA of trypanosome & nematode worm. |
|
|
image of trans-splicing |
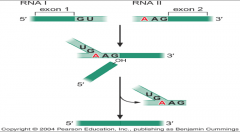
|
|
|
image of singlee gene generate 2 alternate mRNA and thus 2 protein products |
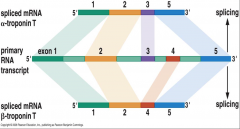
|
|
|
Eukaryotic pre-mRNA processing includes? |
-5' capping -5' structure -5' capping targeting -Polyadenylation
|
|
|
what is 5' capping? |
5' cap is specially altered nucleotide on the 5' end of precursor mRNA & some other primary RNA transcripts in eukaryotes. -process of 5' capping is vital to creating mature mRNA - then able to undergo translation |
|
|
The process of 5' capping generally serves four functions which are? |
1.) regulation of nuclear export 2.) prevention o degradation by exonucleases 3.) promotion of translation 4.) promotion of 5' proximal intron excision |
|
|
what is 5' cap structure? |
5' cap is found on the 5' end of an mRNA molecule & consists of guanine nucleotied connected to the mRNA via an unusual 5' to 5' triphosphate linkage. -Guanosine is methylated on the 7 position directly after capping by a methyl transferase. Referred to as 7-methyl guanosine cap, abbreviatd m7G |
|
|
what is 5' capping targeting? |
- Capping ennzyme complex (CEC) required for 5' capping is -As soon as the 5' end of the new transcription emerges the enzymes transfer to it and begin the capping process.
|
|
|
what is Polyadenylation - Poly A tail? |
linkage of a polyadneylel + mRNA molecule. -most mrNA molecules are polyadenylated at the 3' end of eukary. |
|
|
What're are the 4 functions of Poly A tail? |
1.) protect mRNA from degradation by exonucleases 2.) transcription termination 3.) export of mRNA from the nucleus 4.) mRna translation |
|
|
image of the eukaryotic pre-mrNA processing poly A procress |
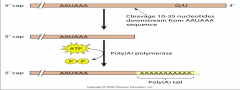
|
|
|
what happens in poly A the process? |
-polyad. occurs during & immediately after transcription of DNA into RNA -After transcription has been terminated, the mRNA chain is cleaved -After mRNA has been cleaved, around 250 adenosine residues are added to the free 3' end at the cleavage site.l -The reaction is catalyzed by polyaden polymerase |
|
|
look at image |
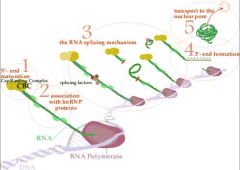
|
|
|
what happens in eukary mRNA export? |
once processed (capped, intron-free & polyaden), MRNA is packaged and exported from the nucleus into the cytoplasm for translation movement from the nucleus to the cytoplasm is an active and carefully regulated processes. -the damaged, misprocesses & liberated introns are retained in the nucleus & degraded. -a typical mature mRNA carries a collection of proteins including SR (serine-argenine rich protein) and another group of proteins binding specifically to exon-exon junctions that id's it absent mrRNA destined for transport |
|
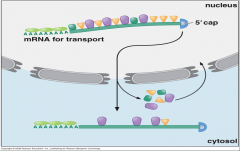
describe the pics. |
export takes places through the nuclear pore complex -once in cytoplams , some proteins are discarded an are then imported back to the nucleus for another cyles of mrNA transport. Some proteins sty on the mana to facilitate translation. |

