![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
20 Cards in this Set
- Front
- Back
|
What are the most blatant and visible aspects of the cell?
|
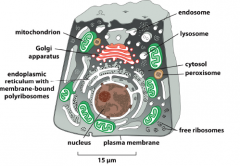
-membranous compartments
|
|
|
How are eukaryotic membranous compartments created, maintained and destroyed? Ie Nucleus and ER
& evidence? |
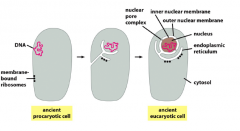
-organelles cannot be created from scratch but from pre-existing cells
-nucleus and ER originated from prokaryotic cell with DNA, and ribosomes. Membrane formed to protect DNA, which extended to become ER with ribosomes attacked to it. -Evidence: inside ER and golgi have same environment as the extracellular space -nuclear bilayer is continous with ER and associated with ribosomes as well |
|
|
What are 3 ways to move molecules b/w compartments?
|
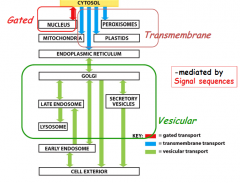
1. Gated transport: RNA leaves nucleus to cytosol and proteins enter nucleus, controlled by selective pore/translocators in nuclear membrane
2. Vesicle Transport: Proteins/Lipids leave from the ER to Golgi and to other compartments (PM/lysomes) 3. Transmembrane transport: proteins from cytosol to ER lumen or mitochondria via membrane-bound protein translocators |
|
|
Basic facts of ER
|
-50% of membrane in a cell is the ER
-makes 10% of total internal volune -site of synthesis of most membrane proteins and secreted proteins -lipid biosynthesis -net-like structure |
|
|
Is rough ER or smooth ER responsible for major site of membrane protein synthesis?
|
-rough ER
|
|
|
T/F: All protein synthesis initiates on functionally identical ribosomes in the same manner?
|
TRUE
|
|
|
How does the mRNA encoding a protein engage ribosomes to the ER? (co-translational)
|
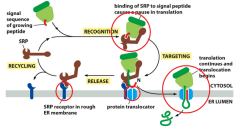
-mRNA has an ER signal sequence which binds to a signal receptor protein. This SRP guides it to a SRP receptor in the ER membrane. SRP receptor will bring it to the protein translocator that will bind to the signal sequence so that translation will continue and translocation will begin.
|
|
|
What does the signal sequence also do besides helping the mRNA encoding a protein bind to the ER membrane? Energy source?
|
It can also serve as a start-transfer signal, by attracting a signal peptidase to cleave the signal peptide on the protein. Once cleaved, the translocator will close and the signal and translocator will be recycled.
-energy source comes from the translating/ribosome machinery |
|
|
How are fully translated proteins transferred/translocated in the ER in EUK and BACTERIA???
|
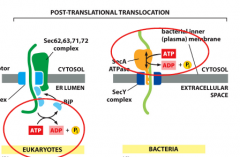
-also called post-translational translocation via active transporters requiring ATP
-EUK: PUll method where ATP is hydrolyzed inside of lumen with BIP. -BAC: push method, where ATP is hydrolyzed in the cytosol (SEC A ATPASE) |
|
|
How does soluble proteins and integral proteins get through the ER?
|
-soluble proteins have start-signal sequence that will be cleaved
-integral proteins may have: -start signal and stop signal -internal signal -internal signal with stop signal |
|
|
How does NH2 end face cytosol?
|
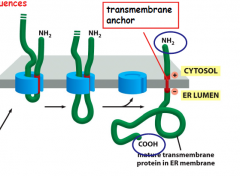
-Basic aa (+) is before the un-cleaved signal peptide
|
|
|
How is a protein with a 2 pass transmembrane constructed?
|
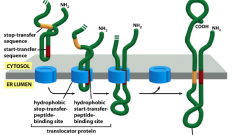
-one un-cleaved start signal sequences and one stop sequence
|
|
|
What are types of post-translational modifications in the ER?
|
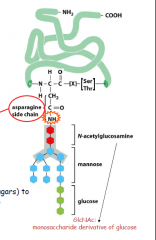
-cleavage of signal peptide
-glycosolations (N-glycosolations of asn side chains) which is adding oligosacchride to the NH grp of asn side chain -sugar trimming if glycoslyated |
|
|
How is a protein glycoslyated in the ER membrane?
|
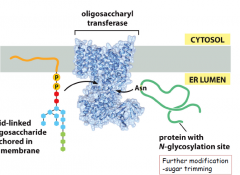
-oligosacchryl transferase will transfer a lipid-linked oligosacchride anchored in the ER membrane to a protein with a N-glycosylation site in the ER lumen
|
|
|
What are the purposes of glycoslyation?
|
1. Promote protein folding
2. protective from proteolytic enzymes 3. signal transduction/cell-cell recognition |
|
|
What is the purpose of N-linked glycoslyation?
|
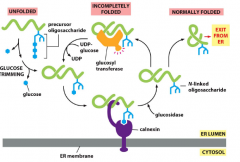
-establish glyco-code that marks progression of protein folding and mediates binding to chaperon proteins
-help to retain incompletely folded proteins in ER lumen |
|
|
What happens to misfolded proteins in the ER?
|
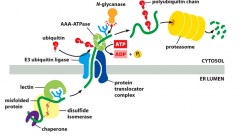
-They are transferred outside of ER to cytoplasm to be degraded. THis is done through intermediate enzymes
1. Unfolding of proteins - disulfide isomerase 2. Forming polyubiquitin chain on protein while exiting ER - protein translocator complex+E3 ubiquitin ligase+ AAA-ATPASE (atp hydrolysis) 3 Unlinking n-glycoslyation - N-glycanase 4. degradation of cell - proteosomes |
|
|
What is the purpose of forming GPI anchored proteins in PM of ER?
|
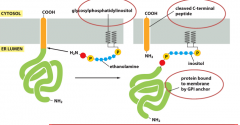
-This permits these proteins to be specifically shed fromPM under certain conditions through phospholipases
|
|
|
Where is the site of lipid synthesis and assembly of lipids?
|
-on cytosolic side of ER where all necessary substrates are found
|
|
|
Scramblase vs Flippase?
|
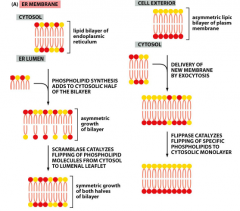
-They are both phospholipid tranferase
-Scrramblase is nonspecific and will make membrane have symmetric growth on both halves of the bilayer by flipping lipids on cytosolic side to ER lumen side - Flippase is head-group specific and will flip lipids so that the lipid composition is assymmetrical in the bilayer |

