![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
29 Cards in this Set
- Front
- Back
|
Topoisomerase I |
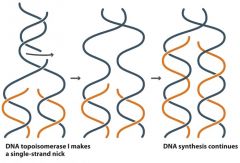
- relieves tension in parent DNA molecule and unwinds strands
|
|
|
DNA Polymerase I |
- Prokaryotic - 1 subunit - 5' → 3' endonuclease activity - 3' → 5' endonuclease activity - involved in replication and repair of DNA - acts on lagging strands, joining Okazaki Fragments
|
|
|
DNA Polymerase III |
- Prokaryotic - at least 10 subunits - 3' → 5' endonuclease activity - main replicating enzyme - synthesises DNA on leading strand |
|
|
DNA Polymerase α |
- Eukaryotic - 4 subunits - No endonuclease activity - Acts as a primer during replication |
|
|
DNA Polymerase δ |
- Eukaryotic - 2 or 3 subunits - 3' → 5' endonuclease activity (proofreading) - Main replicative enzyme |
|
|
Primase |
- Eukaryotic - Prokaryotic - produces RNA based primer
|
|
|
DNA Helicase |
- Breaks H bonds between base pairs - e.g. DnaB in bacterial cells |
|
|
Primosome |
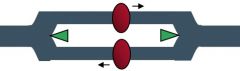
- Formed in E. Coli when primase is attached to the prepriming complex to begin the formation of RNA primers |
|
|
Prepriming Complex |

- formed when DnaB helicase proteins bind to the site of origin - DnaB breaks more base pairs and moves replication fork further from origin |
|
|
DNA-dependent DNA Polymerases |
- Family of enzymes that carry out DNA-dependent DNA synthesis |
|
|
Single Strand Binding Proteins |
- Proteins that protect single strands - Strands may be attacked by nucleases (present to degrade single stranded virus DNA) - Strands may reattach - Eukaryotic example: Replication Protein A |
|
|
DNA Ligase |
- Catalyses the formation of a phosphodiester bond - Joins DNA strands |
|
|
FEN1 |
- Eukaryotic - Cuts 5' end of Okazaki fragments of lagging strand |
|
|
Telomerase |
- Adds TTAGGG (in vertebrates) sequence to 3' end of DNA molecules in telomere regions - Prokaryotic DNA is often circular so does not have ends for telomerase to work on |
|
|
DnaA |
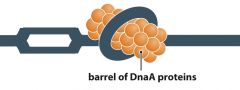
- Prokaryotic - Form 'barrels' of proteins near origin of replication - DNA winds round barrels and breaks base pairs (origin is mostly A-T pairing which is relatively weak compared to C-G)
|
|
|
DnaB |
- Prokaryotic - Helicase enzyme - Breaks more H bonds to move forks from origin - Addition of these form prepriming complex |
|
|
Gamma Complex |
- Prokaryotic - attaches and detaches pol III from the lagging strand during replication - 9-10 subunits |
|
|
Proliferating Cell Nuclear Antigen |
- Eukaryotic - PCNA complex holds Pol δ tightly to DNA - Can be loosened so Pol δ can slide back along strand - 'proliferating': during division, 'Antigen': studied in antibodes |
|
|
Tus Protein |
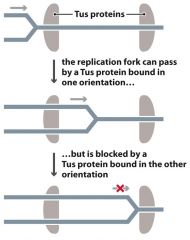
- Part of prokaryotic terminator sequences - allows fork to pass in one direction only |
|
|
Nucleosomes |
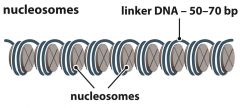
- 'beads' on chromatin - Octomer formed of histones: 2 x H2A, 2 x H2B, 2 x H3, 2 x H4
|
|
|
DNA Glycosylase |
- Prokaryotic - involved in base excision repair - catalyses hydrolysis of n-glycosidic bond between base and sugar - a family of enzymes, each with a specific excision role - leaves an apyrimidinic/purinic (AP) site |
|
|
DNA Polymerase II |
- involved in prokaryotic nucleotide excision repair - low rate of error - can reinitiate DNA synthesis |
|
|
DNA Polymerase γ and DNA Polymerase β |
- Perform base excision repairs in eukaryotic cells |
|
|
Phosphodiesterase + AP endonuclease |
- Prokaryotic - Removes pentose sugar left by removal of base by glycosylase, allowing new nucleotide addition by polymerase and ligase |
|
|
UvrABC Endonuclease |
- Involved in prokaryotic nucleotide excision - moves up and down DNA molecule detecting general damage to helix - uvrA checks for damage, uvrB and uvrC make cuts in molecule, urvB bridges gap in molecule for temporary protection before ligase/polymerase add new nucleotides |
|
|
MutH and MutS |
- Involved in prokaryotic mismatch repair that UvrABC Endonuclease can't detect - MutH detects methylated parent strand - MutS attaches to mismatched base in daughter strand
|
|
|
Ku Proteins |
- Involved in eukaryotic homologous end joining - Ku proteins cap ends of DNA molecule and attract one another, bringing the ends together - Must act quickly or DNA drifts too far apart |
|
|
Histone H1 |
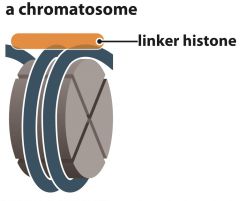
- The linker histone - attaches outside nucleosome |
|
|
Histones |
- Proteins in chromatin - Form nucleosome octomers - links the nucleosome and the DNA strand |

