![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
64 Cards in this Set
- Front
- Back
|
What is a mutation? |
Mutation is a hereditary change in DNA - if it occurs in germlinewill be passed on to offspring |
|
|
Do all organisms carry mutant alleles? |
Every organism carries mutant alleles, whether they manifest indistinct phenotypes or not. |
|
|
When do new mutations occur? |
New mutations can occur in every generation. |
|
|
Why do gene mutations arise? |
Gene mutations occur within individual genes as a result ofchange in nucleotide sequence which can have consequence interms of protein structure or gene expression. |
|
|
What are some causes of mutations? |
Many causes of mutation – DNA replication errors – Spontaneous mutations – Chemicals and irradiation (e.g. ultra violet light) |
|
|
What are somatic mutations? |
Somatic mutations are ones that occur in any cell of the body apart fromgerm cells (egg and sperm). These mutations can not be inherited byoffspring. Somatic mutationscan lead to cancer |
|
|
What are germ-line mutations? |
Germ-line mutations - occur in gametes. These mutations are inheritedby offspring |
|
|
Why are dominant mutations easier to study? |
Dominant mutations are easier to detect than recessive mutations andso are easier to study. |
|
|
Do mutation frequencies differ among organisms? |
Mutation frequencies differ considerably among organisms. |
|
|
Do mutation frequencies vary among genes of a single species? |
Mutation frequencies vary among genes of a single species |
|
|
Do larger genomes have higher mutation frequencies? |
Each species has an average mutation frequency; those with largergenomes have, on average, higher mutation frequencies. |
|
|
Do some genes have higher mutation frequencies and why might this be? |
Mutation frequencies for some genes are considerably higher thanothers in a given genome; genes that have elevated mutationfrequencies are called hotspots of mutation. One reason formutational hotspots is large gene size |
|
|
What is mutation frequency defined as in diploids? |
In diploids mutation frequency is defined as the number of mutationalevents in a given gene over a defined period of time (most often perDNA replication cycle or per cell generation) |
|
|
What are Base-pair substitution mutations? |
the replacement of one nucleotide base pair by another |
|
|
What are transition mutations? |
Transition mutations: one purine replaces another, or one pyrimidinereplaces another. A ↔ G (purine ↔ purine) (A·T ↔ G·C) C ↔ T (pyrimidine ↔ pyrimidine) (C·G ↔ T·A) |
|
|
What are transversion mutations? |
Transversion mutations: a pyrimidine is replaced by a purine or viceversa. |
|
|
What is a silent mutation? |
1. Silent mutation: a base-pair change that does not alter the resultingamino acid sequence due to the redundancy of the genetic code |
|
|
What is a missense mutation? |
Missense mutation: a base-pair change that results in an amino acidchange in the protein |
|
|
What is a nonsense mutation? |
Nonsense mutation: a base-pair change that creates a stop codon inplace of a codon specifying an amino acid |
|
|
Define a synonymous or silent mutation |
Synonymous or silent mutation – changes one codon for an amino acid to another codon forthat amino acid – no change in amino acid (silent mutation) |
|
|
Define missense mutation |
Missense mutation – changes codon for one amino acid to codon for anotheramino acid – also called nonsynonymous mutation – the effect of changing an amino acid will depend on the roleof that amino acid in protein function |
|
|
What is a neutral mutation? |
Missense mutations differ in severity – conservative amino acid substitution substitutes chemicallysimilar amino acid, less likely to alter function often called aneutral mutation e.g. alanine (-CH3) for glycine (-H) |
|
|
What would be the effect of a nonconservative mutation? |
Missense mutations differ in severity – nonconservative amino acid substitution substituteschemically different amino acid, more likely to alter function – consequences for protein depend on the function of theamino acid within the 3D structure e.g. glycine (-H) for glutamate (-CH2-CH2-COO-) |
|
|
Where in the three letter code is more likely to produce a change in amino acid? |
Many amino acids have more than one codon.A mutation in the first letter of the three letter code is morelikely to produce a change in amino acid. |
|
|
Define a nonsense mutation |
Nonsense mutation– change codon for amino acid into translation termination(stop) codon |
|
|
What is the effect of a nonsense mutation? |
Nonsense mutation results in premature termination oftranslation. truncated polypeptides often are nonfunctional |
|
|
What is a frameshift mutation? |
Addition or deletion of one ormore bases changes thereading frame and alters allthe amino acids downstreamof the mutation |
|
|
What are regulatory mutations? |
Some point mutations alter the amount (but not the amino acidsequence) of protein product produced by a gene |
|
|
What do these regulatory mutations affect? |
These regulatory mutations affect regions such as a. Promoters and other regulatory protein binding sites b. Introns c. regions coding for 5′-UTR and 3′-UTR segments on mRNA |
|
|
What are promotor mutations? |
Mutations that alter consensus sequence nucleotides of promoters arecalled promoter mutations |
|
|
What do these interfere with? |
These interfere with efficient transcription initiation. |
|
|
How might the level of transcription be affected? |
Some promoter mutations cause mild to moderate reductions intranscription levels, whereas others may abolish transcription, somemutations can even enhance transcription. |
|
|
What does efficient splicing of introns from pre-mRNA in eukaryotes require? |
In eukaryotes, messenger RNA (mRNA) is transcribed as pre-messengerRNA (pre-mRNA) which is processed to remove intron regions.Efficient splicing of introns from pre-mRNA requires specific sequences atthe intron-exon boundaries. |
|
|
What are splicing mutations? |
Mutations that alter that alter nucleotides required for efficient splicingat intron-exon junctions are called splicing mutations. |
|
|
What can splicing mutations result in? |
These can result in splicing errors and the production of mutantproteins due to the retention of intron sequences in the mature mRNA. |
|
|
What are cryptic splice sites? |
Some base-pair substitution mutations produce new splice sites thatreplace or compete with authentic splice sites during mRNA processing.Leads to new possible forms of protein product. |
|
|
What are forward mutations? |
Forward mutation: converts a wild-type allele to a mutant allele |
|
|
What are reverse mutations? |
Reverse mutations or reversions: convert mutant alleles to wildtypeor near wild-type |
|
|
What is true reversion? |
True reversion: wild-type DNA sequence is restored by a secondmutation within the same codon |
|
|
What is Second-site reversion? |
Occurs by mutation in a different gene and together the two mutationsrestore the organism to wild-type |
|
|
What happens in induced mutation? |
Induced mutation • action of mutagen, an environmental agent that altersnucleotide sequence • process of inducing mutations by mutagens is calledmutagenesis |
|
|
What happens in spontaneous mutations? |
Spontaneous mutation • arise in absence of known mutagen • may be caused by errors in DNA replication • provide “background rate” of mutation • important for evolution |
|
|
What are the 3 types of spontaneous mutation? |
1.DNA replication errors 2.Spontaneous nucleotide base changes – causes mispairing of DNAnucleotides during DNA replication 3.DNA nucleotide lesions – causes mispairing of DNA nucleotides duringDNA replication for example Depurination – removal of purine group Deamination – removal of amino group |
|
|
What happens in DNA replication errors and what normally corrects them? |
DNA polymerase can insert the wrong nucleotide.This is corrected by the ‘proof reading’ 3’ to 5’ exonuclease function of theDNA polymerase 99% of the time. |
|
|
What causes trinucleotide repeat disorders? |
The trinucleotide repeat disorders are caused by DNA polymeraseslipping during DNA replication and increasing the number of trinucleotide repeats within a gene which results in longer stretches of thesame amino acid within the protein. |
|
|
What is an example of a trinucleotide repeat disorder? |
Example - Huntington’s disease (autosomal dominant) - caused byincrease in length of a polyglutamine region in the Huntingtin protein28 CAG repeats - normal 36 or more CAG repeats - disease symptoms |
|
|
What are tautomers? |
DNA nucleotide bases can occasionally convert to alternative structurescalled tautomers, with slight differences in bonding and placement ofhydrogens |
|
|
what can these Spontaneous Nucleotide Base Changes be? |
amino (normal structure for adenine and cytosine) ↔ imino (rare) keto (normal structure for thymine and guanine) ↔ enol (rare) amino is C−NH2 imino is C=NH keto is C=O enol is C−OH |
|
|
What can tautomeric shifts lead to? |
Tautomeric shifts can lead to base-pair mismatches and incorporation ofincorrect bases during replication |
|
|
What is the most common form of replication error? |
tautomeric shifts |
|
|
What causes induced mutations? |
Induced mutations are produced by interactions between DNA andphysical, chemical, or biological agents that generate mutations |
|
|
What are the types of induced mutation? |
1.Nucleotide base analogs 2.Deaminating agents – remove NH2 group3.Alkylating agents - add methy (-CH3) or ethyl group (-CH2-CH3) 4.Hydroxylating agents – add hydroxyl group (-OH) 5.Oxidative agents - addition of oxygen or removal of hydrogen 6.Intercalating agents 7.Radiation induced DNA damage – ultraviolet light, X-rays, gamma rays, cosmic rays |
|
|
What is an example of base analogs? |
Base analogs are similar to nitrogenous bases of DNA, but have alteredpairing properties e.g. 5-bromouracil (5-BU) replaces thymine and causes a AT > GCtransition |
|
|
What is an example of an alkylating agent? |
Alkylating agents modify base structure by adding methyl or ethylgroups which can result in altered base pairing e.g. EMS (ethyl methanesulfonate) |
|
|
What is an example of intercalating agents? |
Intercalating agents are flat, planar molecules intercalate between basepairs distorting the DNA helix which disrupts DNA replication and causesaddition or deletion of nucleotides leading to frameshift mutations. e.g. proflavin, acridine orange, ethidium bromide. |
|
|
What DNA repair mechanisms? |
There are a number of ways organisms try to repair DNA damage usingcorrect base on the damaged DNA strand. |
|
|
What is photoreactivation repair? |
DNA damage repaired byphotoreactivation enzyme (PRE)which cleaves the thymine dimerbond. In E. coli this enzyme is calledphotolyase. |
|
|
What is Base excision repair? |
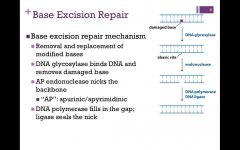
Base excision repair performed bya base specific DNA glycolsylase(in this example uracil DNAglycosylase), AP endonuclease,DNA polymerase and DNA ligase.Uracil is recognised as anoncomplementary base pair andreplaced with the complementarybase (C). |
|
|
What is Nucleotide excision repair? |
Repairs lesions which distort DNA e.g.thymine dimers.In E. coli the UVRA, UVRB and UVRCproteins recognise the lesion and excises13 nucleotides. DNA polymerase I then synthesises thenew DNA strand and DNA ligase joins thesingle stranded nick |
|
|
Human diseases can be caused by mutations in genes that encodeproteins involved in DNA repair mechanisms. What is an example of such a disease? |
Xeroderma pigmentosum - individuals areusually homozygous recessive for a faultyrepair gene. Sufferers are photosensitiveand if their skin is exposed to lightdevelops freckles and patches of intensepigment leading to warty growths that aremalignant and usually fatal. |
|
|
What causes Xeroderma pigmentosum? |
Xeroderma pigmentosum diseasesymptoms are caused by mutant alleles ofsix different genes whose protein productsare involved in DNA repair in humans |
|
|
What can mutations in human genes whose proteins are involved in DNA repair mechanisms cause? |
Mutations in human genes whose proteins are involved in DNA repairmechanisms can cause hereditary cancer. |
|
|
What are some examples of these mutations? |
BRCA2 - hereditary breast cancer MSH2 - Hereditary colorectal cancer MLH1 - Hereditary colorectal cancer |
|
|
What other mutations can cause cancer? |
Somatic mutations in human genes whose proteins are involved in DNArepair mechanisms can cause cancer as well. |

