![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
110 Cards in this Set
- Front
- Back
|
Who do retroviruses infect?
|
Insects --> Fish --> Man
|
|
|
What are the potential infection consequences of retroviruses?
|
- No ill effects
- Tumors - rapid onset or long latency - Wasting diseases, neurological disorders - Immune deficiencies (HIV) |
|
|
What are the genetic properties of retroviruses?
|
- Acquire host cell sequences - oncogenes
- Insert into host cells chromosome - can activate or inactivate genes --> cancer - Rapid genome evolution - acquire mutation through replication and recombination |
|
|
How are viruses classified now?
|
Based on genomic sequence
|
|
|
Enveloped viruses have what extra proteins?
|
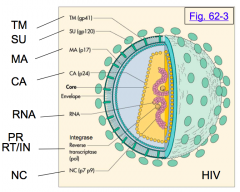
- Env protein embedded in membrane
- Matrix protein under lipid bilayer |
|
|
The capsid (core) of a virus is made of what structural proteins?
|
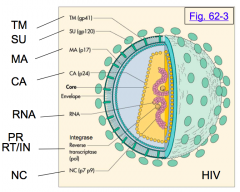
Group specific AntiGens (GAG proteins):
- Matrix proteins (MA) - Capsid proteins (CA) - NucleoCapsid proteins (NC) - Protease (PR) |
|
|
What is the genome of retroviruses?
|
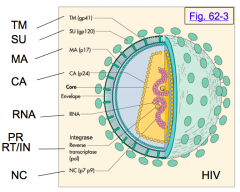
2 copies of a (+) ssRNA genome (only virus that is "diploid" and accounts for recombination potential)
|
|
|
What is the significance of retroviruses having 2 copies of the +ssRNA?
|
- Only virus that is "diploid"
- Accounts for recombination potential |
|
|
What is the simple retrovirus genomic organization?
|
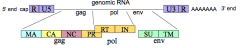
5' end cap
- R (repeat) - U5 (unique to 5' end) - gag gene: MA, CA, NC, PR* - polymerase (pol) gene: RT, IN - envelope (env) gene: SU, TM - U3 (unique to 3' end) - R (repeat) 3' end poly-A tail *PR is in pol reading frame in HIV |
|
|
How do the retroviral genomic RNA get capped (5' end) and polyadenylated (3' end)?
|
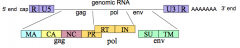
Made by host's Polymerase II - so gets the same modifications as host mRNAs
|
|
|
What proteins does the "gag" gene encode (5'-3')? How are they synthesized?
|
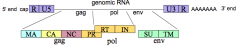
- MA - matrix
- CA - capsid - NC - nucleocapsid - PR - protease* - Made as a polyprotein that then gets clipped (*note - PR is in pol gene reading frame in HIV) |
|
|
What proteins does the "pol" gene encode (5'-3')? How are they synthesized?
|
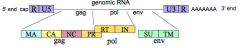
- RT - reverse transcriptase
- IN - integrase - Made as an extended polyprotein * PR is in the reading from of pol gene in HIV (5' end) |
|
|
What proteins does the "env" gene encode (5'-3')? How are they synthesized?
|
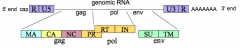
- SU - surface domains
- TM - transmembrane domains - Made as a precursor and gets cleaved into SU an dTM domains |
|
|
What is the complex retrovirus genomic organization (e.g., HIV)?
|
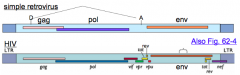
- Organized similarly to similar retroviruses (w/ gag, pol, env genes)
- Except numerous additional genes/proteins - mRNAs for additional genes ("accessory proteins") are generated by complex alternative splicing |
|
|
What is the point of the accessory proteins of complex retroviruses (HIV)?
|
Endow HIV with its unique properties, various functions
|
|
|
What are the two phases of the replication cycle in retroviruses?
|
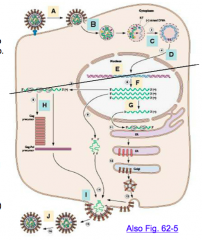
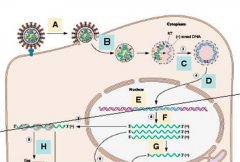
A-D - precedes integration
E - integration into host DNA F-J - post-integration |
|
|
What are the steps of replication of retroviruses?
|
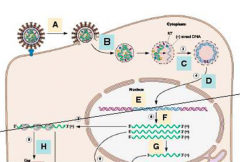
A - adsorption
B - penetration and uncoating C - reverse transcription D - transit into nucleus E - integration into host DNA F - viral RNA synthesis, host pol II G - RNA processing H - virion protein synthesis I - assembly and budding J - capsid maturation (proteolysis) |
|
|
What happens during the first step of the replication cycle of retroviruses?
|
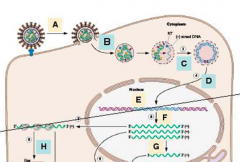
A - adsorption
- Virus binds cell via env protein and a host cell receptor - HIV receptor is CD4 / CCR5 - Other viruses use amino acid transporters, LDL-like receptor, etc. |
|
|
What happens during the second step (after adsorption) of the replication cycle of retroviruses?
|
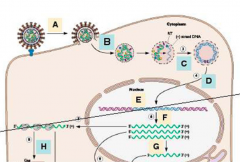
B - penetration
- Viral envelope fuses with cell membrane either at cell surface or in endosomes after endocytosis |
|
|
What happens during the second step (after penetration) of the replication cycle of retroviruses?
|
B - uncoating
- Genomic RNA is only partially uncoated (remains in protein particle in cytoplasm - prevents translation) - RT, IN and some of the gag proteins remain associated with incoming genomic RNA - Proteins are needed to convert ssRNA genome to dsDNA, nuclear import, and integration |
|
|
What is the purpose of the genomic RNA of retroviruses only be partially uncoated after penetration into the host cell?
|
Do not want translation at this point - protein particle prevents association with ribosomes - goal is to convert ssRNA to dsDNA
|
|
|
What happens during the third step (after partial uncoating) of the replication cycle of retroviruses?
|
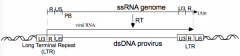
C - reverse transcription
- Conversion of ssRNA to dsDNA w/ RT - Integrated DNA ("Provirus") is longer than the template RNA w/ U3 and U5 duplicated at ends to form Long Terminal Repeat (LTR) |
|
|
What are the functions of Reverse Transcriptase (RT)?
|
- RNA-dependent DNA polymerase (copies RNA --> DNA)
- DNA-dependent DNA polymerase (copies a 2nd strand of DNA from first strand) - Error-prone polymerase, ~5 errors made per genome, leads to rapid evolution and drug resistance |
|
|
What happens during the fourth and fifth steps (after reverse transcription) of the replication cycle of retroviruses?
|
D/E - Transit to nucleus & Integration
- Integration requires dsDNA have access to host DNA - Integrase recognizes and is specific for sequences at ends of dsDNA (i.e., ends of U3 and U5) - Integration reaction is at random part of host sequence via IN protein - Now, virus is a permanent resident of host cell's DNA |
|
|
How do retrovirus' dsDNA gain access to host DNA for integration?
|
- Many retroviruses can't cross nuclear membrane and need cell division to integrate
- HIV CAN cross nuclear membrane, important for infection of non-dividing cells, and for gene therapy vectors |
|
|
Where does integration of retrovirus' dsDNA into host DNA occur?
|
Essentially random - integration reaction is NOT specific for host sequences
|
|
|
What happens during the sixth step (after integration of dsDNA into host) of the replication cycle of retroviruses?
|
F - proviral transcription
- LTR directs synthesis of viral RNA - 5' U3 binds transcription factors - Beging transcription at R region |
|
|
How do the LTR (Long Terminal Repeats) direct synthesis of viral RNA?
|
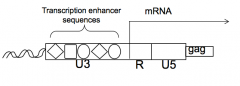
- 5' U3 contains binding sites for cellular transcription factors required for high level RNA synthesis
- U3 has signals recognized by the cell's transcription machinery, which directs transcription at beginning of R region |
|
|
How does U3 on the LTR influence which tissue/cells a retrovirus is active in (e.g., tropism)?
|
- Spectrum of proteins that bind the U3 determines which tissues/cells it is active in
- E.g., HIV LTR requires transcription factor NFκB (only expressed in activated T cells) |
|
|
How does HIV RNA regulate which types of cells it is transcribed in?
|
- HIV LTR requires transcription factor NFκB, which is only expressed in activated T cells
- Not transcribed in infected memory T cells because memory cells do not express NFκB |
|
|
What happens during the seventh step (after proviral transcription) of the replication cycle of retroviruses?
|
G - RNA processing
- All viral RNAs are polyadenylated, some must be spliced to generate the env mRNA - A large portion must remain full length to serve as gag-pol mRNA and as genome for progeny virions - Complex retroviruses (HIV) do much more alternative splicing |
|
|
What are the three fates of retroviral RNA:
|
1. Full length RNA --> genomic RNA
2. Full length RNA --> gag-pol mRNA 3. RNA splicing --> env mRNA (and others) |
|
|
Why is retroviral RNA splicing incomplete?
|
To preserve full-length RNA for genomic RNA and gag-pol mRNA
|
|
|
What happens during the eighth step (after RNA processing) of the replication cycle of retroviruses?
|
H - Translation
- Most abundant protein is gag and gag-pol - Gag initiates at an AUG start codon, and ends at a stop codon at end of gag - Gag-pol is made from same AUG start coon, but ribosomes ignore or circumvent the gag stop codon and continue to end of pol (only happens about 5% of time, so RT and IN proteins are less abundant, also not needed in surplus, than gag) - Both gag and gag-pol proteins are eventually cleaved by protease (PR) domain to release individual proteins - Env protein is made from spliced mRNA on rER --> golgi --> inserted into plasma membrane |
|
|
How are the gag and gag-pol proteins related?
|
- Both start at the same AUG start codon
- gag protein stops synthesis at stop codon - gag-pol protein ignores gag stop codon and continues to end of pol (only happens 5% of time) - Eventually cleaved by protease (PR) |
|
|
What are the env proteins?
|
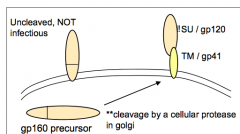
- Env gp160 precursor protein
- Cleaved into gp120 and gp41 by cellular protease - Cleavage must happen because gp160 cannot support membrane fusion (virus would be made but could not fuse w/ target cell) |
|
|
What happens during the ninth/tenth steps (after translation) of the replication cycle of retroviruses?
|
I/J - virion assembly and budding
- Packaging requires signal Psi (ψ) contained in unspliced RNA; splicing removes ψ signal - Budding - viral gag and gag-pol polyproteins recruit RNA and assemble under cell surface; gag protein interacts w/ env and budding occurs as particle forms - Maturation - proteolysis of gag and gag-pol by PR occurs after budding, causing protein rearrangements and core to become more dense - Viral particles still form and bud if proteolysis is inhibited, but viruses are not infectious |
|
|
What is the action of PR inhibitors?
|
- Inhibit proteolysis of gag and gag-pol by PR (which occurs after budding)
- Viral particles still form and bud if proteolysis is inhibited but they are NOT INFECTIOUS |
|
|
How were retroviruses discovered?
|
As agents isolated from naturally occurring tumors in animals, that, when inoculated into naïve animals, would again cause tumors
|
|
|
What are the types of retroviruses?
|
- Non-transforming = non-acute or slow tumor viruses
- Transforming = acute tumor viruses |
|
|
What are the characteristics of non-transforming retroviruses?
|
- Non-acute or slow tumor viruses
- Tumors take 6 mo to 1 yr to appear - Do not transform cells in culture (low freq., i.e., no cancer phenotype) - Viruses do not contain oncogenes - Tumors are caused by activation or inactivation of host genes |
|
|
What are the characteristics of transforming retroviruses?
|
- Acute tumor viruses
- Tumors occur within weeks - Causes tissue culture cells to become "transformed" or cancer-like - Viruses harbor a mutated copy of a cellular gene involved in growth control, "oncogene" --> rapid tumor onset upon introduction via infection |
|
|
What was the first oncogene to be identified?
|
Src - in Rous sarcoma virus (avian virus)
|
|
|
Which groups of people are more commonly infected by HIV?
|
- Homosexual men
- Heroin addicts - Hemophiliacs - Haitians |
|
|
HIV patients have a decline in what cells?
|
CD4 T cells
|
|
|
When was the death rate from AIDS highest for males under 40 years old? What caused a decline?
|
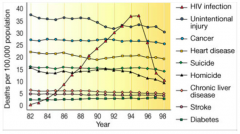
- Peaked in 1994-1995
- Declined w/ use of HAART (Highly Active Antri-Retroviral Therapy) |
|
|
In 2011, how many adults and children are living with HIV?
|
34 million
|
|
|
What are the known transmission routes of HIV infection?
|
* Sexual transmission (vaginal and anal)
- Perinatal transmission (intrauterine, breast milk) - can be reduced w/ antiviral treatment - Inoculation in blood (transfusion, needling sharing, needle-stick) |
|
|
What are some of the additional genes/proteins in the complex retrovirus, HIV?
|
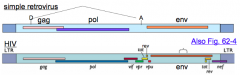
- 6 Accessory proteins (Tat, Rev, Vif, Vpu, Vpr, Nef)
- 2 Regulatory proteins (Tat and Rev) - 2 Restriction factors (Vif and Vpu) |
|
|
What are the 6 additional accessory proteins of HIV required for?
|
Replication and / or pathogenesis
|
|
|
What is the function of Tat and Rev? Type of proteins?
|
- Essential, crucial for viral replication (attractive targets for therapy)
- Regulatory proteins |
|
|
What kind of protein is Tat? Function?
|
- Regulatory protein / accessory protein of HIV
- Tat = Transactivator of Transcription - Absolutely required for transcription (mRNA synthesis) |
|
|
Which protein is absolutely required for transcription?
|
Tat
|
|
|
What kind of protein is Rev? Function?
|
- Regulatory protein / accessory protein of HIV
- Rev = Regulator of Virion Expression - Allows structural gene expression by promoting transport of unspliced RNA from nucleus to cytoplasm (export) |
|
|
Which protein allows structural gene expression by promoting transport of unspliced RNA from nucleus to cytoplasm?
|
Rev
|
|
|
What is the function of Vif and Vpu? Type of proteins?
|
- Viral proteins that overcome cellular defenses, or "restrictions"
- Restriction factors |
|
|
What kind of protein is Vif? Function?
|
- Restriction factor / accessory protein
- Vif = Virion Infectivity Factor - Causes a cellular antiviral protein (deoxycytidine deaminase) to ge degraded; otherwise is incorporated into new virions where block RT in next cell by inducing massive mutations in viral dsDNA |
|
|
What protein causes deoxycytidine deaminase to be degraded?
|
Vif
|
|
|
What kind of protein is Vpu? Function?
|
- Restriction Factor / Accessory protein
- Promotes virion release from cells by inhibiting a host protein "tetherin" which otherwise blocks release of virus from the cell - Works on other enveloped viruses |
|
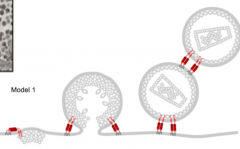
What protein promotes virion release from cells by inhibiting "tetherin" (red linkages)
|
Vpu
|
|
|
Are Tat, Rev, Vif, and Vpu good anti-viral targets?
|
Hypothetically yes, but so far nothing has been developed that targets them effectively
|
|
|
What receptor/cells does HIV bind to?
|
CD4 on immune cells:
- CD4 Helper T-cells - Dendritic Cells - Macrophages - Microglia |
|
|
How does HIV bind to / affect CD4 Helper T-cells?
|
Binds; this is the main cell population that is depleted in AIDS
|
|
|
How does HIV bind to / affect Dendritic Cells?
|
Can bind HIV at CD4 receptor, but can not be productively infected; can assist in dissemination
|
|
|
How does HIV bind to / affect Macrophages?
|
Infected via binding to CD4 receptors, but not efficiently killed - reservoir of virus production
|
|
|
How does HIV bind to / affect Microglia?
|
Binds via CD4 receptors; leads to brain infection - contributes to AIDS dementia
|
|
|
For HIV what is necessary for membrane fusion?
|
CD4 binding AND co-receptor
|
|
|
What are the HIV tropisms / co-receptors? Types of cells that are infected?
|
- M-tropic (R5-tropic) & CCR5 co-receptor - infects primary T-cells and macrophages (NOT T-cell lines)
- T-tropic (X4-tropic) & CXCR4 co-receptor - infects primary T-cells and T-cell lines (NOT macrophages) |
|
|
What is M-tropic binding responsible for? What kind of people have this kind of infection?
|
Initial infection and transmission, and predominates in asymptomatic persons
|
|
|
What is T-tropic binding responsible for? What kind of people have this kind of infection?
|
Associated with disease progression, arises at AIDS stage of infection
|
|
|
What chemokines can specifically inhibit M-tropic HIV? How?
|
- RANTES, MIP-1α, MIP-1β
- Occupy receptor (CCR5 co-receptor) |
|
|
What chemokines can specifically inhibit T-tropic HIV? How?
|
- Cytokine Stromal Derived Factor 1 (SDF-1)
- Natural ligand for CXCR4 |
|
|
What do HIV co-receptor expression levels correlate with?
|
Cell permissiveness
|
|
|
What is the basis for strain tropisms?
|
Env sequence of different HIV types have preference for different co-rereceptors (most concern is for M-tropic virus, which is source of person-to-person transmission)
|
|
|
What is the explanation for why some rare individuals remain seronegative despite high-risk behavior and presumable repeated viral exposure?
|
- For some, explanation is a 32bp-deletion in CCR5 gene (co-receptor for M-tropic HIV)
- Causes non-functional CCR5 - WT:Δ32 heterozygotes (~10%) get infected but progress to disease more slowly - express about half as mucha s normal - Δ32:Δ32 homozygotes (~1%) are highly resistant to infection - normal despite lack of CCR5 expression on surface |
|
|
What are the steps of the fusion process of HIV?
|
1/2: Env initially contacts CD4 and induces a conformation change in Env to expose co-receptor binding site
3/4: gp41 "fusion domains" are exposed, and fusion domain enters cell membrane 5: co-receptor (CCR5) engagement triggers a "snapback" of N- and C- terminal helical regions of gp41 (yellow and red cylinders), which brings membranes together and fuses them |
|
|
What happens in the first two steps of the the fusion process of HIV?
|
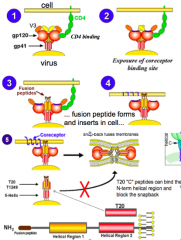
1/2: Env initially contacts CD4 and induces a conformation change in Env to expose co-receptor binding site
|
|
|
What happens in the third and fourth steps (after initial contact of Env w/ CD4) of the the fusion process of HIV?
|
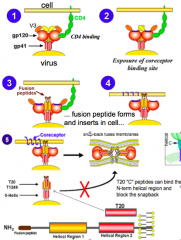
3/4: gp41 "fusion domains" are exposed, and fusion domain enters cell membrane
|
|
|
What happens in the fifth step (after gp41 fusion domain is exposed) of the the fusion process of HIV?
|
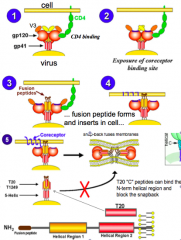
5: co-receptor (CCR5) engagement triggers a "snapback" of N- and C- terminal helical regions of gp41 (yellow and red cylinders), which brings membranes together and fuses them
|
|
|
What can block snapback in the 5th step of HIV fusion process?
|
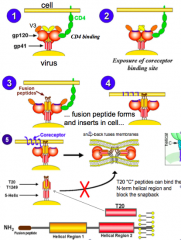
T20 "C" peptides (Fuzeon antiviral) can bind the N-terminal helical region and block snapback
|
|
|
What are the steps of pathogenesis to HIV infection?
|
1. Initial HIV infection
2. Spread to lymph nodes 3. Virus infects T cells, replicates, and gets into circulation (viremia) 4. Asymptomatic phase |
|
|
Where/how does HIV infection usually occur?
|
At mucosal surfaces or by blood products
|
|
|
After an initial HIV infection, what happens?
|
Spreads to lymph nodes - DC cells can bind and carry HIV to nodes, where T cells reside and are infected
|
|
|
What happens after HIV spreads to the lymph nodes?
|
Virus infects T cells, replicates to high levels, and spills into circulation (viremia)
|
|
|
What happens during the asymptomatic phase of HIV?
|
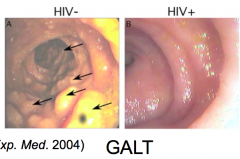
- FDC traps virus, keeps viremia low, but nodes (especially GALT - gut associated lymphoid tissue) are major sites of replication (1 billion/day)
- GALT deteriorates late in infection (destruction of lymph tissue) |
|
|
What is the most important site of infection of HIV?
|
GALT - gut associated lymphoid tissue
|
|
|
What are the disease mechanisms of HIV?
|
- Direct killing of CD4 T-cells by HIV
- Indirect effects on infected CD4 T cells - Impairment of immune system function |
|
|
How does HIV directly kill CD4 T cells?
|
- Massive virus production leads to membrane linkage and death
- Syncytia (fused cells) induced by fusion of infected cell with uninfected cells (via Env on infected cell interacting w/ CD4/CCR5 on uninfected cell) - cells eventually die (could kill uninfected cells via by-stander effect) - Apoptosis induced by infection, some evidence that cells undergo apoptosis even if unproductive infection |
|
|
How does HIV indirectly kill CD4 T cells?
|
- Immune response kills infected cells, important for clearing initial viremia
- Soluble gp120 may bind uninfected cells, now susceptible to ADCC (antibody dependent cellular cytotoxicity) |
|
|
How does HIV impair immune system function?
|
- CD4 T-cell function altered, and loss of CD4 T cell help --> leads to severely compromised immune system
- Infected macrophages are dysfunctional --> aberrant immune system |
|
|
What are the clinical features of the acute phase of HIV infection and seroconversion?
|
- Initial burst of virus production coincides w/ decreased CD4 T-cells
- Early vigorous CTL, subsequent humoral response, w/ FDC help, clears viremia - Immune response only "appears" to control infection - High level virus production persists in lymph nodes / GALT |
|
|
What are the clinical features of the asymptomatic phase of HIV infection?
|
- Continued strong immune response, but gradual decline in CD4 counts
- Progression measured by CD4 counts, CD4:CD8 ratio, "viral load" by measuring RNA by PCR; (patients w/ low "set-point" have better prognosis) |
|
|
What are the clinical features of the symptomatic/AIDS phase of HIV infection?
|
- Late in infection, CD4 cells depleted (< 200 / µL)
- Immune system begins to fail - Viremia increases - Patients susceptible to many opportunistic infections |
|
|
How does the amount of virus change throughout the three stages of an HIV infection (acute / asymptomatic / symptomatic-AIDS)?
|
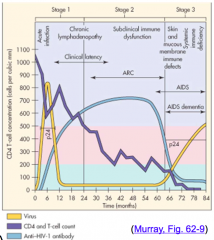
- Stage 1 - acute - Initial high amount of virus
- Decline during clinical latency through Stage 2 - asymptomatic - Stage 3 - symptomatic/AIDS - amount of virus rises (YELLOW) |
|
|
How does the CD4 and T-cell count change throughout the three stages of an HIV infection (acute / asymptomatic / symptomatic-AIDS)?
|
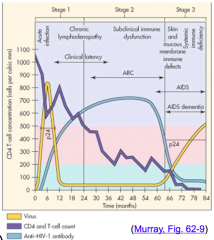
Continually gets lower through all three stages
(PURPLE) |
|
|
How does the amount of anti-HIV-1 antibody change throughout the three stages of an HIV infection (acute / asymptomatic / symptomatic-AIDS)?
|
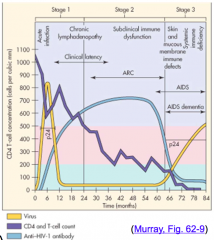
- Stage 1 - increases
- Stage 2 - plateaus and declines at end (beginning of AIDS) - Stage 3 - very low (BLUE) |
|
|
When are tests done for HIV?
|
- Identify infected persons to initiate treatment
- Identify "carriers" who may transmit HIV to others - Follow course of disease - Evaluate efficacy of treatment |
|
|
What serological laboratory tests are done to diagnose HIV?
|
- Ab ELISA - initial screening, detect Ab to virus (rapid oral test - inexpensive w/ fast results)
- Ag ELISA - detect p24 antigen earlier than Ab - Western Blot - confirmation test, uses patients Ab to detect HIV protein |
|
|
What laboratory tests are used to diagnose HIV?
|
- Serological (Ab ELISA, Ag ELISA, Western Blot)
- RNA RT-PCR |
|
|
How does RT-PCR diagnose HIV? Uses?
|
- Detects virus in blood
- Quantitate virus in blood - Very sensitive (≤ 50 HIV RNA copies / mL) - Detects virus before seroconversion (high risk groups, newborns) - Gauges viral load in asymptomatic patients w/ low titers (important for prognosis) |
|
|
What are the therapeutic targets for HIV?
|
- Any step in replication cycle, especially virus specific proteins
- RT inhibitor - Protease inhibitors - Fusion inhibitor (T-20) - Entry inhibitors (CCR5 co-receptor antagonist) - Integrase inhibitor - HAART - Highly Active Anti-Retroviral Therapy |
|
|
Is it necessary to completely block HIV replication to remain healthy?
|
No - progression is related to viral load
|
|
|
Which kind of HIV therapy can reduce viral load by 30-100x alone?
|
Protease inhibitors
|
|
|
What is HAART? Outcomes?
|
- Highly Active Anti-Retroviral Therapy
- Triple therapy ("cocktails") of different inhibitors - Virtually eliminate virus production in some individuals for many years - Virus undetectable in plasma, increased CD4 cell counts and obvious clinical benefit - Long-term patients experience toxicity |
|
|
What did HAART allow determination of?
|
HIV replication dynamics, 1/2 life of HIV
|
|
|
How long does it take to clear free virus and infected T cells w/ HAART?
|
~2 months
|
|
|
How long does it take to clear other "compartments w/ longer half-lives (macrophages and FDCs) w/ HAART?
|
~1-2 years
|
|
|
How long does it take to clear infected memory T cells w/ HAART?
|
Long-lived, can detect virus from those on HAART for > 5 years (resting T cells); would need > 75 years to clear this compartment
|
|
|
Why is the enthusiasm for HAART and a "functional cure" for HIV tempered?
|
- Not all patients respond to HAART
- Drug regimen is difficult to follow (getting easier w/ 1x/day pills) - Toxic effects seen in long term HAART users - Inaccessible pool of virus |
|
|
What kind of drugs are being developed to treat HIV?
|
- New attachment inhibitors (anti-CD4 antibody)
- New CXCR4 and "dual receptor" inhibitors, and anti-CCR5 antibody - RT inhibitors to common drug-resistant RT viruses - New integrase inhibitors - Maturation inhibitors that work on gag and gag-pol proteins |

