![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
293 Cards in this Set
- Front
- Back
|
Why does water have a high melting and boiling point? |
Hydrogen bonds must be broken to melt ice and vaporize water. |
|
|
Heat of Vaporization of water?
Shape of water? |
Heat of Vaporization: 40 kJ/mol (high heat of vaporization)
Shape: tetrahedryl (104.5) |
|
|
On average, each water molecule is bonded to ____ other water molecules. In ice, water forms ____ hydrogen bonds, increasing it's volume. |
Average: 3.4 other water molecules Ice: 4 hydrogen bonds
This is why ice floats-- because its density is decreased by the increase in volume. |
|
|
Hydrogen Bond Characteristics |
|
|
|
When is a hydrogen bond strongest? |
When three atoms lie in a straight line |
|
|
Water competes with molecules for hydrogen bonds. The strength of hydrogen bonds can be increased by ________. |
Shielding the bond from the water molecules
e.g., double helix DNA where base-to-base hydrogen bonds are within helix and not exposed to water molecules. |
|
|
DNA is ______ charged. Histones are ______ charged.
They're attracted to one another in water, however the level of attraction is dependent on _______ concentration. |
DNA = Negative Histones = Positive
Depends on salt concentration. At high concentrations, there are more ions between DNA and histones, decreasing their attraction to one another. |
|
|
Amphipathic molecules |
Have a polar and nonpolar region |
|
|
Hydrophobic Effect |
Non-polar molecules forced to cluster.
|
|
|
Ionization state of a group is determined by ______ as well as ______. |
1. Dissociation constant 2. pH of solution |
|
|
Dissociation Constant of Water |
10^-14
It is small, so only a small fraction of molecule is dissociated |
|
|
Ionization State of Weak Acid and how it relates to pH |
|
|
|
Because pH is a logarithmic scale:
|
1. 90% protonated 2. 10% protonated |
|
|
Example of a strong acid and the pH
HCL --> H+ + Cl- |
The concentration of [H+] = 0.1M.
So the pH = -log(0.1) = 1, and the solution is acidic. |
|
|
Equation to determine the pH |
pH = -log [H] |
|
|
Common structure of all amino acids? |
Central carbon (alpha) bound to carboxyl and amino group.
H2N--C--CO2H
|
|
|
At a pH of 7, the amino group and the carboxyl group of an amino acid are both ______. |
Ionized
It's called a Zwitterion |
|
|
All proteins are optically active because it has a central carbon with four different groups.
What is the exception? |
Glycine is the only AA that is not an optical isomer because it has an H in place of the R group. |
|
|
Are amino acids L- or D- optical isomers? |
All AAs are L-Amino Acids |
|
|
Which are the non-polar alipathic Amino Acids? (7) |
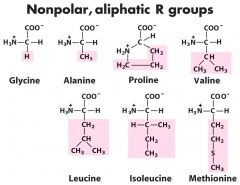
|
|
|
What is the unique characteristic of proline? |
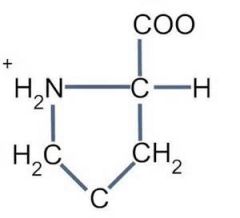
1. Stiff and can change direction of a peptide chain.
2. Only AA with secondary amine on it that folds back on itself (all others have primary amine group).
- So the amino and carboxyl side are on one side and cannot rotate in free space like other AAs.
|
|
|
Two Amino Acids with Hydroxyl Groups? |
1. Serine: essentially an alanine with an OH
2. Threonine |
|
|
Three aromatic amino acids? |
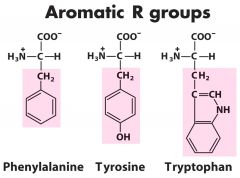
1. Phenylalanine (alanine w/ phenyl group) 2. Tyrosine (OH on end of the phenyl group) 3. Tryptophan (endol group) |
|
|
What determines the UV absorption of proteins? |
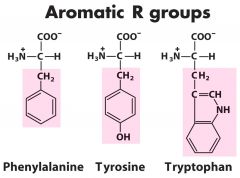
Aromatic amino acids |
|
|
Intrinsic fluorescence of proteins is mostly due to _______. |
Tryptophan |
|
|
Positively charged R groups? (3) (BASIC) |
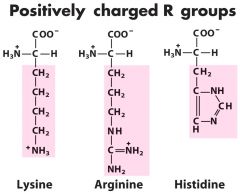
1. Lysine 2. Arginine 3. Histidine
|
|
|
Positive charged amino acids are acidic or basic? |
POSITIVE = BASIC
NEGATIVE = ACIDIC |
|
|
Negative charged amino acids are acidic or basic? |
POSITIVE = BASIC
Negative = Acidic |
|
|
Negatively Charged Amino Acids (2)? (Acidic) |
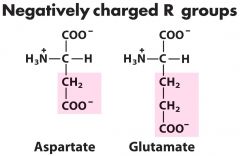
1. Aspartate (aspartic acid w/ H) 2. Glutamate (glutaminc acid w/ H) |
|
|
What two amino acids have an amide side chain? |
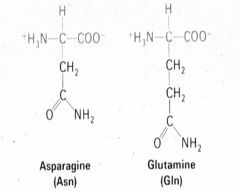
1. Asparginine (Asn) 2. Glutamine (Gln) |
|
|
What two amino acids contain sulfur? |
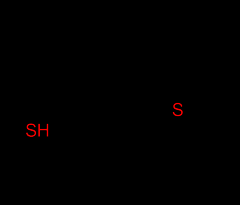
1. Cysteine (Cys) - very unstrable - spontaneously oxidizes to disulfide bonds
2. Methionine (Met)
|
|
|
Unique characteristics of Cysteine? |
1. Unstable 2. Spontaneously OXIDIZES to for disulfide bonds |
|
|
3 Modified Amino Acids |
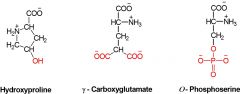
1. Hydroxyproline 2. ɤ - Carboxyglutamate 3. O-Phosphoserine |
|
|
In a protein, only the AA side chain can ionize. Why? |
Because the carboxyl and amine are involved in the peptide bond. |
|
|
Peptide bone is formed by ______. |
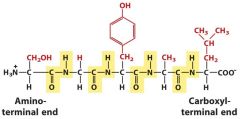
Removal of water between a carboxyl and amino group. It's between the carboxyl carbon of one group and the amine of another.
|
|
|
Typical AA Composition of Proteins |
|
|
|
Characteristic of Peptide Bone |
1. Partial double bond characteristic
2. Planar because carbon-nitrogen bond has partial double bond character.
3. Rigid and does not rotate around the N-C bond. |
|
|
Alpha helix protein structure is formed by? |
Rigidity of the peptide bond and hydrogen bonding is used to predict alpha helical structure.
|
|
|
Beta sheet protein structure |
Hydrogen bonding and peptide bond rigidity used to predict B-sheet structure
|
|
|
Difference between physical properties of alpha helix and beta sheet? |
Alpha helix: can be stretched and absorb water
Beta sheet: cannot be stretched and does not absorb water |
|
|
Almost all proteins contain some _____ and _____ structure. |
Alpha helix and beta sheet structure |
|
|
Proline and Glycine residues often occur at bends in a peptide chain. Why? |
1. Proline forces the turn 2. Glycine is flexible due to small side chain. |
|
|
Conjugated proteins |
Contain group other than amino acids (e.g., lipoprotein contains lipid group) |
|
|
Four levels of protein structure: |
Primary: Sequence of AA and disulfide bridges
Secondary: Steric relationship of AA which are close to one another (e.g., B-sheet, A-helix)
Tertiary: Steric relationship of entire chain (e.g., folding)
Quaternary: number of subunits and geometry of their packing
|
|
|
In a protein (e.g., myoglobin) with many alpha helices, hydrophobic side chains occur where ________. |
The helices are in contact. |
|
|
Types of structural patterns in proteins? (4) |
|
|
|
Structures used to denature proteins? (3) |
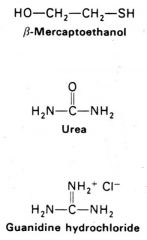
1. Beta-mercaptoethanol 2. Urea 3. Guanidine hydrochloride |
|
|
Proteins can refold spontaneously according to their ______. |
Amino acid sequence.
Some proteins cannot refold (e.g., if there are post-translational modifications) |
|
|
Immunoglobulins on protein function and structure |
1. Specificity coded in AA sequence 2. IgG has quaternary structure and domains 3. Domains have different functions 4. Domains have same structure, evolution by gene duplication |
|
|
In contrast to most proteins, IgG ________. |
Does not have a unique amino acid sequence.
Sequence is variable in N-terminal of heavy and light chains. |
|
|
The variable amino acid sequence occurs in the ______ of the heavy and light chains of IgG. |
First domain of the heavy and light chains
The hyper-variable regions provide binding specificity. |
|
|
Operon |
Unit of DNA containing a cluster of genes under the control of a single promoter. These genes are co-transcribed into an mRNA strand.
Genes in an operon are expressed all together or not at all. |
|
|
Lac Operon (E. coli) and it's regions |
An example of an inducible set of genes. It has three genes in the operon. The regions, from left to right are:
|
|
|
Difference in DNA location for prokaryote and eukaryote |
Prokaryote doesn't have nucleus, so DNA is free floating.
Eukaryote has nucleus, so DNA is separate. |
|
|
Gene expression in prokaryotes is coupled with translation |
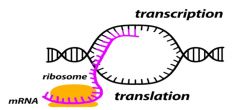
Transcription is tightly coupled with translation
Two Regulation Points: 2. Attenuation: start transcribing but stop before elongation
|
|
|
Prokaryotic gene structure |
1. Core promoter: -35 to -10 region UP element 2. Promoter: DNA sequence around site of transcriptional initiation involved in binding of RNA Polymerase 3. Operator: DNA element that binds specific repressor tightly and prevents transcription 4. Activator binding sites 5. Structural Genes: usually transcribed together in a POLYCISTRONIC mRNA (encodes several proteins from one RNA molecule) |
|
|
Positive and Negative Regulation of Transcription |
POSITIVE: Induction--activator facilitates transcription initiation.
NEGATIVE: Repression--repressor inhibits transcription initiation.
|
|
|
Three structural genes of the Lac Operon |
1. LacZ: B-galactosidase breaks down lactose into glucose and galactose
2. LacY: Permase increases uptake of lactose from outside of cell
3. LacA: Transacetylase
|
|
|
Negative Regulation of Lac Operon |
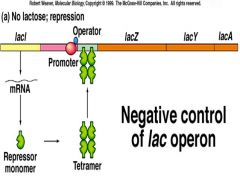
NO Lactose present
Note: NO absolute turnoff for the ZYA genes because LacL is not always repressing at the operon. |
|
|
Depression occurs in the presence of lactose (the inducer) |
If lactose is present, the Lac Operon is working at low levels but needs glucose to have high levels of activity.
|
|
|
Catabolic Repression
If lactose and glucose is present, the cell will ______ before the Lac operon is turned on. |
Cell will use all glucose before Lac operon is turned on. |
|
|
What molecule signals the presence or absence of glucose in the cell for the Lac Operon? |
cAMP.
When glucose is high, cAMP is low.
The Lac promoter is weak. Without CRP-cAMP binding the lacZYA promoter is weak. |
|
|
Positive Regulation of Lac Operon Catabolite Repression |
If glucose present, catabolite repression prevents transcription even if the inducer (e.g., lactose) is present.
|
|
|
Summary of Lactose Operon |
![BOTH NEGATIVE AND POSITIVE TOGETHER (with lactose and no glucose)
1. Lactose is converted to -> allolactose by β-galactosidase, repressor dissociates from operator (transcription increases 20-fold)
2. Without glucose [cAMP] increase...](https://images.cram.com/images/upload-flashcards/71/40/22/7714022_m.png)
BOTH NEGATIVE AND POSITIVE TOGETHER (with lactose and no glucose)
1. Lactose is converted to -> allolactose by β-galactosidase, repressor dissociates from operator (transcription increases 20-fold)
2. Without glucose [cAMP] increases in the cells, cAMP binds to CRP
3. cAMP-CRP binds to CRP site in lac promoter and further stimulates transcription about 50 fold to achieve 1000 fold in total.
|
|
|
Regulatory Schemes of Prokaryote Gene Regulation |
Anabolic Pathway: turned on by substance it breaks down
Catabolic Pathway: turned off by product
|
|
|
Tryptophan Characteristics |
|
|
|
Difference between Tryptophan and Lac Operons |
Lac Operon: inducible
Tryptophan Operon: repressible
|
|
|
Two ways the Tryptophan Operon is regulated |
1. Negative Regulation: high Trp binds repressor and stops transcription.
2. Attenuation: inhibition of elongation
|
|
|
Explain Attentuation in Trp Operon 1. High [Trp] 2. Low [Trp] |
High [Trp] Level:
Low [Trp] Levels:
|
|
|
Difference between Trp repressor (TrpR) and Lac repressor (LacL) |
Trp binds to TrpR and makes it active. Now it can bind to operator and repress. It's normally off.
Allolactose binds LacL repressor and makes it inactive. It's normally on. |
|
|
Structure of the Trp Operon |
TrpR: repressor (unlinked to TrpE-A and has own promoter)
TrpL: leader sequence. Required for attenuation
P: Promoter for RNA Pol
O: Operator: Binding site for trpR
TrpE-A: structural genes
|
|
|
Eukaryotic Gene Expression is Regulated Largely at the Transcriptional Level through what mechanisms (5)? |
REMEMBER: Gene transcription (mostly initiation) is the predominant regulatory mechanism. |
|
|
Effect of euchromatin and heterochromatin on transcription? |
Heterochromatin: Tightly wound and coiled. Inhibitory because transcription factors (TF) cannot gain access to DNA sequence and trigger transcription initiation.
Euchromatin: open and accessible to TF. This allows transcription to occur. |
|
|
Nucleosomes |
Composed of octomers of histones (4 dimers) |
|
|
How is heterochromatin turned into euchromatin? |
1. SWI/SNF factor is an ATPase that drives dissociation of nucleosomes, opening up the heterochromatin. |
|
|
Structure of SWI/SNF Complex |
Has a CENTRAL ATPase subunit in the complex. This can be one of two proteins (but not both):
SWI/SNF complex integrates:
|
|
|
Post-translational modification of histones and two types of epigenetic marks?
|
A histone PTM can be repressive or activating for chromatin structure--depends. These are called EPIGENETIC MARKS. They occur in the AMINO-TERMINAL TAIL.
Lysines: can be acetylated or methylated Serines and threonines: phosphorylated or ubiquinated
REPRESSION MARK: tri-methylation of lysine #9 on histone #3 (H3K9Me3)
ACTIVATION MARK: acetylation of lysine #8 on histone 3 (H3K8Ac)
|
|
|
DNA is negatively charged and histones are positively charged. This is how they stay together.
How does acetylation effect this? |
Acetylation of LYSINES block positive charge, disrupting interaction with negative phosphate backbone of DNA and promotes euchromatin.
Histone Acetylations (HACs) add acetyl groups
Histone Deacetylations (HDACs) remove acetyl groups. |
|
|
Difference between transcription factor and coregulator? |
Transcription Factor: bind directly to DNA at specific sequences.
Coregulators: DO NOT bind to DNA
Both regulate transcription |
|
|
Examples of Transcription Factors (from the eukaryotic gene regulation lecture)
Tata-Binding Protein (TBP |
TBP binds to TATAA box and serves as landing pad for RNA Pol II next to transcription initiation site. |
|
|
Examples of Transcription Factors (from the eukaryotic gene regulation lecture)
AP-1 Protein "TRE Response Element" |
Binds to consensus sequence TGATCA.
AP-1 is a heterodimer of protooncogene products FOS and JUN. It targets genes by:
|
|
|
Examples of Transcription Factors (from the eukaryotic gene regulation lecture)
Vitamin D Receptor (VDR) Vitamin D Response Element (VDRE) |
Heterodimer of Vitamin D Receptor (VDR) and Retinoid-X-Receptor (VDR/RXR) is a transcription factor.
It is activated by the ligand Vitamin D (Calcitriol). |
|
|
Eukaryotic RNA Polymerase II Promoter:
Elements can function both independently and cooperatively. |
Independently
Cooperatively: transcription factors form more highly active complexes when simultaneously bound to adjacent response elements. |
|
|
Four Superclass of Transcription Factors 1. TF Class 2. Examples |
Based on their DNA-binding motifs
Superclass I: Leucine Zipper Factors (bZIP)
Superclass II: Zinc-Coordinating DNA Binding Domain
Superclass III: Helix-Turn-Helix
Superclass IV: Beta-Scaffold Factors with Minor Groove Contacts
|
|
|
Superclass I Leucine Zipper (bZIP) |
Two subunits. Hydrophobic interactions between to Leucines hold then together and form "zipper". The basic region interacts with DNA.
Examples:
AP-1 is a heterodimer of FOS and JUN proteins. AP-1 binds TREs (TPA response element). TPA activates phosphorylation cascade, which ends by activating JUN.
|
|
|
AP-1 is a transcriptional effector of MAPK signaling. How does this work? |
Mitogens/Growth factors lead to activation of AP-1 and transcription of TRE-containing genes (e.g., c-fos gene) by initiating the mitrogen-activate protein kinase (MAPK) signaling cascade.
1. Induction of c-fos expression via transcription ERK is the MAPK that phosphorylates TCF, which with SRF and EVI-1, activates c-fos transcription.
2. Newly synthesized FOS protein combines with JUN, followed by phosphorylation of dimer by same cascade, binding to AP-1 elements (TREs)--induction of transcription. |
|
|
Superclass II Zinc Coordinating DNA Binding Domain Nuclear Receptors |
Nuclear receptors (NRs) are prototypes of Zn-Coordinating DNA Binding TFs. Can be heterodimer or homodimer.
All heterodimer NRs have common RXR unit, with other subunit providing unique activity.
NRs (and all other TFs) have two functional domains:
Most NRs also have a Ligand-Binding Domain that induces a conformational change that opens the DNA-binding interface of the molecule.
Activating domains in NRs serve as interaction sites for other transcription activating proteins that form the bridge to RNA pol II at the start.
|
|
|
Zinc Finger Motifs |
Zn confers alpha helices (with connecting beta strands) that insert into the MAJOR groove of DNA for nucleotide-specific binding.
All members of SUPERCLASS 2 use this basic Zn finger motif for DNA binding. |
|
|
Superclass III Heliz-Turn-Helix Transcription Factors |
Includes Homeodomain TFs (HOX), Forkhead TFs (FOX), and ETS factors.
Helix-turn-helix motif provides DNA binding interface of 3 alpha helices that bind to major groove of DNA. |
|
|
Superclass IV Beta-Scaffold Factors w/ Minor Groove Contacts |
Examples: p53 tumor suppressor, STAT family
Note, they contact the MINOR and major groove of the DNA.
|
|
|
What are the two obligatory domains of all transcription factors (TFs)? |
1. DNA binding domain 2. Transcription-activating domain |
|
|
Nuclear Receptors (NRs) use recruitment of _______ to promote assembly and activation of the RNA Pol II initiation complex. |
Mediator complex.
Mediator is last part of the bridge. It extends from regulatory DNA element to the RNA Pol II.
|
|
|
Binding of an activating transcription factor (TF) to its target DNA sequence elicits three kinds of transcription-activating machinery. They are? |
1. SWI/SNF 2. HAT complexes 3. Mediator |
|
|
In eukaryotes, _____ domain is the kinase that phosphorylates the C-terminus of Pol II. This is the last step in activating the Pol II to begin transcription. |
CDK-8 domain |
|
|
Alpha Helix |
Peptide chain wrapped into a helix and hydrogen bonded to itself. |
|
|
Beta Sheet |
Peptide chains are in extended line conformations. Adjacent chains are hydrogen bonded to each other. |
|
|
Collagen Helix |
Linear helix created by three peptide chains wrapped around each other. |
|
|
Beta Turns |
Conformations adapted when a peptide chain has to sharply change directions in a folded protein. |
|
|
A zymogen is |
Self-inhibiting |
|
|
Enzyme Kinetics Pre-Steady State Burst |
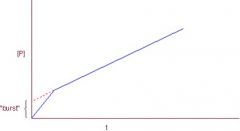
Intermediate lasts on order of milliseconds of seconds. Initial burst is evidence of an ACYL INTERMEDIATE in the reaction
Presteady state burst continues until product to enzyme ratio is 1:1 and then it begins a steady state reaction. When [P]/[E] > 1, RXN is in steady state.
|
|
|
Serine Protease contain a CATALYTIC TRIAD comprised of what three amino acids? |
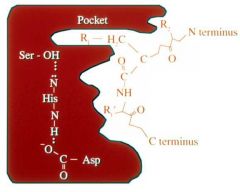
Serine Histidine Aspartate |
|
|
Active site of an enzyme has residues important for what? |
1. Chemistry of RXN 2. Binding of substrate |
|
|
Binding Residues in R1 and R2 position for: 1. Trypsin 2. Chymotrypsin 3. Thrombin |
Trypsin: R1: K, R R2: --
Chymotrypsin: R1: F, W, Y R2: --
Thrombin: R1: R R2: P
|
|
|
Drug inhibitor design is based on _______.
Meaning of a
|
Shape complimentarity.
Low Ki: only a little bit of drug needed to get 50% inhibition (GOOD)
High Ki: takes a lot of drug to get to 50% inhibition (BAD) |
|
|
Steps to make an inhibitor protein (3) |
Low Ki: keep it High Ki: discard |
|
|
A common combination of amino acids that should be included in drug design? |
1. Proline 2. Arginine
"it didn't take a computer to figure that out" |
|
|
Thrombin PPACK |
PPACK
Catalytic Triad:
|
|
|
A peptide bond cannot be hydrolyzed without the help of _____. |
An enzyme (e.g., Serine Protease) |
|
|
Hydrolysis of a peptide bone by serine protease |
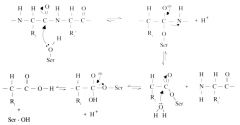
1. Hydrolysis of peptide bond 2. Hydrolysis of bond with serine protease, releasing the enzyme.
Enzymes must always be released--not used up during reaction. |
|
|
Summary of how an enzyme helps get from substrate to product. |
![Enzyme must be complementary to transition state to work. If enzyme is complementary to substrate, transition state cannot form and RXN won't occur.
The order of the reaction is:
Enzyme [E] + Substrate [S]
Enzyme-Substrate [ES] Complex
Tr...](https://images.cram.com/images/upload-flashcards/71/88/67/7718867_m.jpg)
Enzyme must be complementary to transition state to work. If enzyme is complementary to substrate, transition state cannot form and RXN won't occur.
The order of the reaction is:
Transition state stabilization:
|
|
|
Difference in Delta G (free energy) for catalyze vs. uncatalyzed reaction |
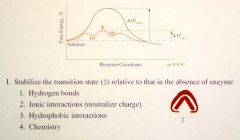
Delta G for uncatalyzed reaction: much higher
Delta G for catalyzed reaction: low |
|
|
What can proteins do to stabilize the transition state (4)? |
1. Make H+ bonds 2. Ionic interactions (neutralize charge) 3. Hydrophobic interactions 4. Chemistry |
|
|
Define an acid and base |
ACID:
BASE:
|
|
|
Linear order of catalytic triad |
Serine-->histidine-->aspartate |
|
|
General base catalysis |
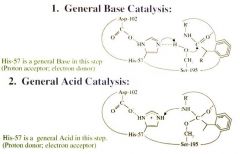
General Base Catalysis
|
|
|
What is the pKa of:
|
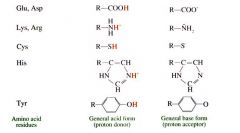
Glu, Asp: 4.5
His: 6.5
|
|
|
Steps of Serine Protease Function (8) |
|
|
|
Serine Protease Function: Step 1 General Base Catalysis |
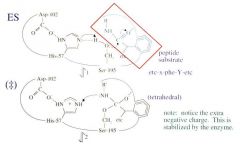
The shape and charge of the transition state is different than the shape and charge of the ES.
Peptide bond shape: planar Transition state shape: tetrahedral
The two dashed lines represent two hydrogen bonds that stabilize the extra negative charge in the transition state. This is worth 4 orders of magnitude.
|
|
|
Serine Protease Function: Step 2 General Acid Catalysis |
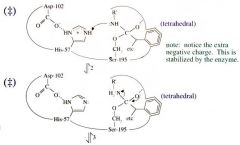
Proton is added to the N of the serine, making it (+) charged.
The (-) charge of the oxygen is now removed and the bond between the carbon and the positively charged N is broken |
|
|
Serine Protease Function: Step 3 Rearrangement of Breaking of Peptide Bond
|
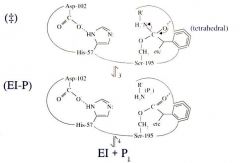
Rearrangement of breaking of the peptide bond
|
|
|
Serine Protease Function: Step 4 Product #1 is Released |
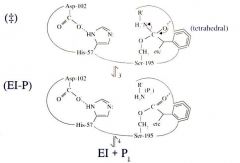
Product 1 (P1) is released
|
|
|
Serine Protease Function: Step 5 Water Enters Active Site (Hydrolysis) |
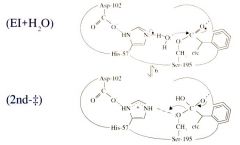
Water enters the active site (hydrolysis)
|
|
|
Serine Protease Function: Step 6 Start of Deacylation |
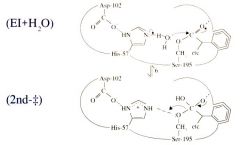
Start of Deacylation
|
|
|
Serine Protease Function: Step 7 General Acid Catalysis (His-57) and Breaking of the Acyl Enzyme Intermediate Bond to Ser-195 |
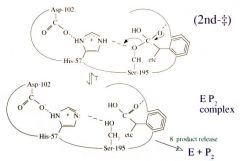
General acid catalysis (His-57) and breaking of the acyl enzyme intermediate bond to Ser-195
|
|
|
Serine Protease Function: Step 8 Product #2 is Released |
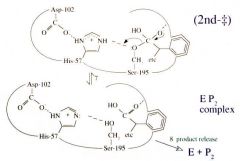
Product #2 is released. |
|
|
Serine Protease Transition State Analogues (2) |
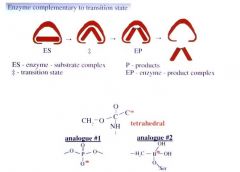
Bind 2-3 times orders of magnitude better than substrate because enzyme wants to entice/stabilize transition state.
(Tetrahedral negatively charged shape)
|
|
|
Parts of a Free Energy Diagram |
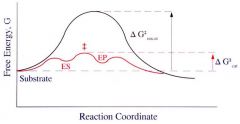
Note, delta Gcat is much lower than delta G uncat.
Also note location of the transition state. |
|
|
First Rate law for first and second order |
First Order Rate Law: (s^-1) v = k[S]
Second Order Rate Law: (M^-1s^-1) v = k [S1][S2] |
|
|
Units of first and second order rate laws? |
First order: s^-1
Second order: M^-1s^-1 |
|
|
First order equation |

h: Plancks' constant K: Boltzmann constant T: Temperature R: Gas constant
Units: s^-1
*Lowe the delta G, higher the rate |
|
|
In first order equation, how does delta G and the rate (k) relate? |
Lower the delta G, higher the rate (k) |
|
|
What does rate (k) in enzyme kinetics represent? |
How many micromolar of substrate can get transformed per minute. |
|
|
Reasons why you want to get an assay as soon as possible? |
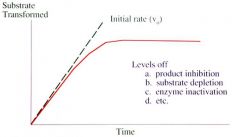
They level off:
|
|
|
Methods for measuring enzyme rate? (6) |
|
|
|
You can only use the Michaelis-Menten equation to get Vmax if _________. |
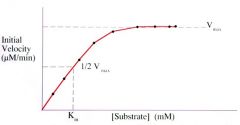
If the graph is a hyperbolic curve. |
|
|
Units of Km (@ 1/2 Vmax) and units of Vmax? |
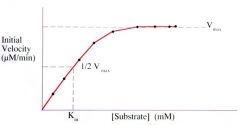
Km (@ 1/2 Vmax): = Millimolar (mM) (note this is the substrate concentration)
Vmax = Micromolar/minute (uM/min) (note this is the rate) |
|
|
Be careful, doesn't equal 1/2 Vmax because the units wouldn't be correct. |
![Km is the [substrate] that gives 1/2 Vmax.](https://images.cram.com/images/upload-flashcards/72/00/10/7720010_m.jpg)
Km is the [substrate] that gives 1/2 Vmax.
|
|
|
Dependence of Initial Velocity (Vo) on [Substrate] |
![Note, at low [S] we can eliminate [S] from the equation because it's negligible.
Note, at high [S] we can eliminate Km from the equation because it's negligible.
Now, Vo = Vmax (the two [S] cancel out)](https://images.cram.com/images/upload-flashcards/72/00/58/7720058_m.jpg)
Note, at low [S] we can eliminate [S] from the equation because it's negligible.
Note, at high [S] we can eliminate Km from the equation because it's negligible.
|
|
|
Using a graph to determine Vo and Vmax |
![Note, at Vo the substrate is essentially zero, so [S] is eliminated from the bottom of the equation.
You get Vo from the data.
At high [substrate], note that Vo = Vmax](https://images.cram.com/images/upload-flashcards/72/00/64/7720064_m.jpg)
Note, at Vo the substrate is essentially zero, so [S] is eliminated from the bottom of the equation.
You get Vo from the data.
At high [substrate], note that Vo = Vmax |
|
|
Steady State Kinetics |
![Rate of ES formation is equal to the rate of ES breakdown
[S]total >> [E]total and [S] = [S]total
So the change in ES over time is zero
Waiting for product to form (k2 or kcat) is really slow.](https://images.cram.com/images/upload-flashcards/72/00/79/7720079_m.gif)
Rate of ES formation is equal to the rate of ES breakdown
As soon as S leaves the ES complex, a new S immediately comes into the spot.
|
|
|
Remember Km is the substrate concentration to give 1/2 Vmax. It is not always the binding constant. |
Remember Km is the substrate concentration to give 1/2 Vmax. It is not always the binding constant. |
|
|
Association constant and dissociation constant using k values. |
k1/k-1 = association constant
k-1/k1 = dissociation constant |
|
|
Initial rate (Vo) is proportional to the rate limiting step (k2) and the amount of substrate present (ES) |
Vo = K2 [ES]
Initiate rate (Vo) is proportional to rate limiting step (k2 or kcat) and the amount of substrate present [ES]
|
|
|
Memorize this equation:
Vmax = k2 [Et]
k2 = rate limiting step [Et] total enzyme concentration |
Vmax = k2 [Et]
So if you know Vmax and k2, you'll be able to determine the amount of [Et] in the blood.
k2 is found experimentally |
|
|
When you flip over a hyperbolic curve, you get a _________.
This changes the Michaelis-Menten equation used with the hyperbolic curve to the _______, which is used with new graph. |
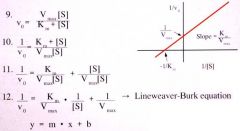
Double Reciprocal Plot (straight line)
Lineweaver-Burk Equation |
|
|
Michaelis-Menten Equation |
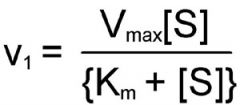
V1 should really be Vo. |
|
|
Lineweaver Burk Equation |
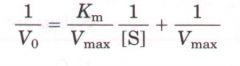
This is in the form of Y= mx + b
Slope: Km/Vmax
y-int: 1/Vmax
x-int: 1/km |
|
|
Comparing high and low km values |
Low km = HIGH Affinity
High km = LOW affinity
|
|
|
General Equation for MM and enzyme kinetics |
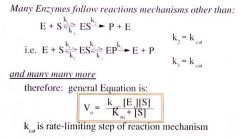

You can get Kcat, Km, and Vmax from steady state kinetics |
|
|
Not all enzymes follow the Michaelis-Menten Enzyne Steady State Kinetics.
Anything that's not a perfect hyperbolic curve cannot use MM equation |
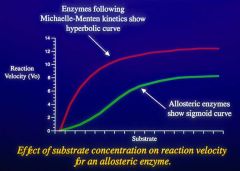
Allosteric enzymes show a sigmoid curve (or anything other than hyperbolic)
It can bind via either a cooperative process or anticooperative process |
|
|
Allosteric enzymes |
Heterotropic modulator
Homotropic modulator:
|
|
|
Types of inhibition (2) |
1. Irreversible inhibition (suicide substrates)
2. Reversible Inhibition
Inhibitors can have linear, hyperbolic, or parabolic effects |
|
|
Competitive Inhibitor on Lineweaver Burke Plot |
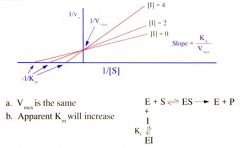
MORE SUBSTRATE has an effect
|
|
|
Non-competitive Inhibitor on Lineweaver Burke Plot |
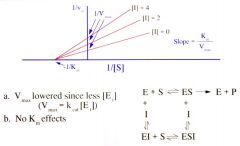
|
|
|
Non-Competitive Inhibitor Site vs. Allosteric Binding |
Non-Competitive Inhibitor Site:
Allosteric Binding:
|
|
|
Uncompetitive Inhibitor on Lineweaver-Burke Plot |
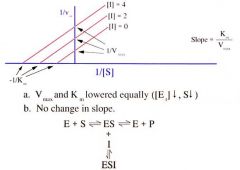

Inhibitor binds ONLY to the ES complex. It cannot bind in the absence of substrate. |
|
|
On Lineweaver Burke Plot, how do you find Vmax? |
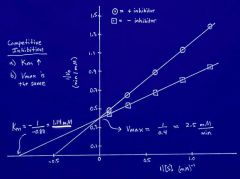
Do 1/Y-intercept. This is the same as 1/Vmax. |
|
|
Equation to determine concentration from spectroscopy? |
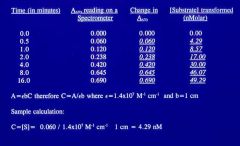
A = εbC or C = A/εb
where, ε = 1.4 x 10^7 M^-1 cm^-1 b = 1 cm
C is the concentration of substrate |
|
|
Impact of genetic disorders |
3-7% Dx with genetic disorder
|
|
|
Define:
|
Gene: functional and physical unit of heredity passed from parent to offspring - hereditary unit: molecularly a sequence of DNA required for the production of a functional product.
Locus: position of a gene on a chromosome
Mutation: change in DNA sequence - permanent, heritable |
|
|
Types of Mutations (outcomes) (4) |
1. Loss of function: reduction or loss of Fx - often recessive phenotypes
2. Gain of function: increase in NORMAL Fx - often dominant phenotypes
3. Novel property: confers new property/Fx - often cancer, rearrangement
4. Dominant-negative allele: disrupts Fx of a wild type allele in same cell - frequently dominant phenotypes |
|
|
Information about human genome |
Genome: all DNA in organism or cell
|
|
|
Define:
|
Allele: alternative form of genetic information at a particular locus
Polymorphism: at least two relatively common alleles at a locus (e.g., blood type) |
|
|
Define:
|
Genotype: individuals genetic makeup--his or her DNA sequence at given locus
Phenotype: OBSERVABLE expression of a genotype |
|
|
Pedigree symbols
|
Twins Shown as a carrot If line connects carrot, they're IDENTICAL
Divorce Double lines through connection
Miscarriage Triangle
Unknown Sex Diamond |
|
|
Single Gene Traits/Disorders |
Traits that are determined by alleles at a single locus. |
|
|
Define:
|
Dominant: phenotypically expressed in heterozygotes
Recessive: phenotypically expressed only in homozygotes for mutant allele
Codominant: alleles that are both expressed when they occur together (e.g., ABO blood groups) |
|
|
Define:
|
Homozygous: identical alleles at a given locus
Heterozygous: different alleles at a given locus
Compound heterozygote: two different mutant alleles at a given locus |
|
|
Degree of relationships: 1st, 2nd, and 3rd degree |
1st Degree: parents, siblings, offspring
2nd Degree: grandparents, aunts, nieces
3rd Degree: first cousins, etc. |
|
|
Pedigree Pattern of Autosomal Dominant Inheritance |
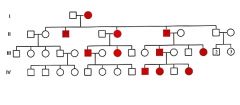
|
|
|
Factors complicating pedigree analysis:
|
New mutation: frequent cause for affected individual with no family history (AD)
Germline Mosaicism: mutation in a germline in one parent
Penetrance: proportion of individuals with a disease genotype who express the disease phenotype. If it's not 100%, it's reduced.
Variable Expressivity: severity of disease may vary greatly. Extent of expression of phenotype. |
|
|
Factors complicating pedigree analysis:
|
Pleiotropy: when a gene has multiple, seemingly different unrelated, phenotypic effects
Genetic Heterogeneity: similar phenotype caused by different genotypes
Linkage: co-inheritance of two or more non-allelic genes at nearby loci |
|
|
Homozygous Autosomal Dominant Genotype |
Typically much more severe than heterozygote
Consider in rare situation where two heterozygotes mate |
|
|
Examples of genetic disorders with delayed age of onset |
Huntington disease Myotonic dystrophy Familial Alzheimer disease AD breast CA AD Parkinson disease |
|
|
Examples of Autosomal Dominant Conditions |
Marfan syndrome Achondroplasia (dwarfism) Familial (early) Alzheimer Disease Huntington Disease Familiar Hypercholesterolemia Familial Breast CA (BRCA 1 or 2) |
|
|
Pedigree pattern of Autosomal Recessive Inheritance |
|
|
|
Examples of Autosomal Recessive Conditions |
Sickle cell disease Cystic fibrosis Tay-Sachs disease Hemochromatosis Phenylketonuria (PKU) Thalassemias |
|
|
X-Linked Recessive Inheritance Pattern |
|
|
|
Lyon Hypothesis |
One X chromosome in each somatic cell is randomly inactivated early in embryonic development.
Ensures dosage compensation |
|
|
X-Linked Recessive Example Disorders |
Duchenne muscular dystrophy Hemophilia A Color blindness Glucose-6-phosphate dehydrogenase (G6PD) deficiency |
|
|
X-Linked Dominant Inheritance Pattern |
|
|
|
X-Linked Dominant Condition Example |
Hypophosphatemic (Vit-D resistant) Rickets
Incontinentia Pigmenti Type I |
|
|
Other factors to consider in genetic patterns
|
Sex-limited traits: autosomal genes expressed in only one sex
Sex-influenced traits: autosomal genes where the same genotype is expressed in different frequencies, depending on sex
Phenocopy: phenotype produced by environmental factors that mimics a genetically determined trait. |
|
|
Define:
|
Polygenic: Inheritance of a trait determined by many genes at different loci--each with a small additive affect.
Multifactorial: inheritance of a trait determined by combination of genetic and environmental factors. Complex inheritance. |
|
|
Qualitative vs. Quantitative Trait |
Qualitative: individual either has it or they don't (e.g., pyloric stenosis)
Quantitative: trait with measurable quantity that differs among individuals.
|
|
|
Threshold Model |
You either have it or you don't (Yes or No)
|
|
|
Multifactorial (Complex) Inheritance |
Cause most common adult conditions and congenital malformations
|
|
|
Recurrence risk in Multifactorial (Complex) Inheritance |
Risk to 1st degree relative: square root of population risk
Recurrence risk is higher if > 1 family member affected.
|
|
|
Examples of multifactorial malformations |
|
|
|
Empiric risk |
Based on observational data from many studies
|
|
|
Probability Calculations "and" vs. "or" |
And: probability of two independent events (A and B) both occurring. Multiply.
p(A) x p(B) e.g., 1/2 x 1/2 = 0.25 (1/4) -------------------------------------------------------------
Or: probability of either of two mutually exclusive events occurring. Addition.
p(A) + p(B) e.g., 1/4 + 1/4 = 0.5 (1/2) |
|
|
Hardy-Weinberg Law (and both equations) |
In absence of influences, allele and genotype frequency will remain constant from one generation to the next.
p + q = 1 p^2 + 2pq + q^2 = 1
|
|
|
Hardy-Weinberg Law Summary of Steps (4) |
1. Determine genotype frequency (count people) e.g., 100 people: 64 are AA; 32 are Aa; 4 are aa
2. Count genes (remember, two genes per person)
3. Calculate gene frequency
4. Calculate genotype frequency
|
|
|
Hardy-Weinberg Principle Assumptions (5) |
|
|
|
Genetic Drift |
Chance fluctuation of allele frequencies over time in a small population |
|
|
Founder effect |
In small population, distribution of genes can be determined by founders, the people at the top of the pedigrees that started the kindred or population. |
|
|
Gene flow |
Slow diffusion of genes across a barrier (physical or cultural) |
|
|
Define
|
Assortative Mating: non-random mating based on phenotype
Fitness: measured by number of an individual's offspring; depends on differences in survival and fertility. |
|
|
Fitness differences in Dominant vs. Recessive |
|
|
|
Autosomal Recessive Disease Hardy-Weinberg Information |
RULE: frequency of mutant allele (q) is the square root of the disease incidence (q^2)
e.g., disease incidence (q^2) is 1/40,000
RULE: carrier frequency is 2q (really 2pq, but p=1) |
|
|
Autosomal Dominant Disease Hardy-Weinberg Information |
RULE: frequency of mutant allele is about 1/2 the incidence of the trait.
e.g., If disease incidence is 1/500
1/500 = 2pq = 2p and p = 1/1000 |
|
|
X-Linked Recessive Disorders: Hardy-Weinberg Information |
Males are hemizygotes (XY), so disease frequency equals gene frequency
Females have two genes (XX), so Hardy-Weinberg can be used to calculate carrier frequency.
|
|
|
X-Linked Dominant Disorders: Hardy Weinberg Information: |
Incidence of affected males = q
Incidence of affected females = 2(p)(q)
|
|
|
Bayes Analysis |
|
|
|
Define genetic terms:
|
Presymptomatic: eventual development of disorder is certain if mutation present
Predisposition: eventual development of symptoms is likely, but not certain, in presence of mutation. |
|
|
Breast Cancer |
|
|
|
BRCA1 and BRCA2 Lifetime Cancer Risks |
BRCA1 Mutation
BRCA2 Mutation
Both have risk for prostate, pancreatic, and male breast CA. |
|
|
Most common genetic cause of mental retardation and birth defects? |
Chromosome abnormality |
|
|
Each DNA molecule has been packaged into a mitotic chromosome that's ______ fold shorter than it's length. |
10,000 |
|
|
Chromosome Abnormalities
|
Constitutional: present at birth, usually at conception of shortly thereafter. Leading cause of pregnancy loss and retardation.
Acquired: develop in somatic cells; associated with cancers. Can be solid or fluid.
|
|
|
Mammalian Cell Cycle Interphase |
During interphase, DNA is decondensed. Cell spends significant amount of time here. (n = haploid chromosome; c = # of chromatids)
G1 Phase: interval between mitosis and replication
S Phase: DNA replication 7 hours
G2 Phase: interval between S and mitosis
|
|
|
Steps in Mitosis PMAT |
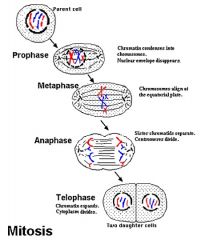
Prophase:
Metaphase:
Anaphase:
Telophase:
|
|
|
Three types of chromosome structures (3) |
1. Metacentric 2. Submetacentric 3. Acrocentric |
|
|
Metacentric Chromosome |
Two sister chromatids connected in middle by a centromere.
|
|
|
Submetacentric Chromosome |
Centromere that is closer to one side than the other.
This is how most human chromosomes look.
Short arm = p Long arm = q (think, "petite") |
|
|
Acrocentric Chromosome |
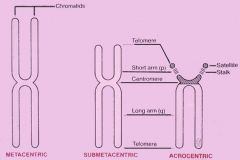
Have a unique short arm side.
|
|
|
What are the acrocentric chromosome (5)? |
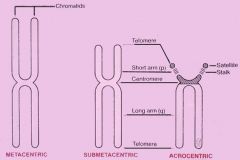
Chromosome #: 13-15, 21, and 22 |
|
|
Short arm of acrocentric chromosomes don't have _________.
They do have _________ that code for ______. These are called _______. |
DO NOT have coding genes.
They do have repetitive sequences to code ribosomal RNA.
These are called Nuclei Organizing Regions. |
|
|
Mitotic Nondisjunction |
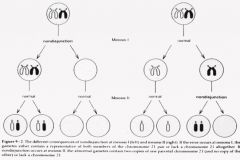
Mitotic error that results in: trisomy, monosomy, and normal chromosome distribution.
3 Cell Lines:
|
|
|
Mosaicism |
Coexcistence of two different cell lines
Trisomy cell line (from non-disjunction) coexists with the normal mitotic cell (disomy)
|
|
|
Meiosis |
Comprised of ONE ROUND of DNA replication but TWO cell divisions (M1 and M2).
|
|
|
Meiosis I |
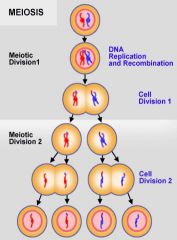
Two chromosomes (maternal and paternal) become two cells with one chromosome (2n-->1n).
|
|
|
Meiosis II |
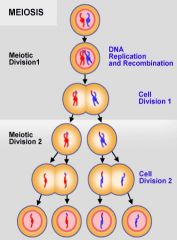
Two cells with one chromosome each comprised of two sister chromatids split into four cells
Note: there is no DNA replication in 2nd meiotic phase. |
|
|
Meiotic Nondisjunction Meiosis I |
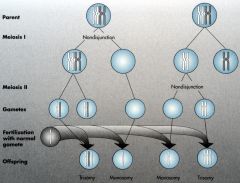
If it occurs during 1st cell division, one normal and one without any chromosomes.
|
|
|
Meiotic Nondisjunction Meiosis II |
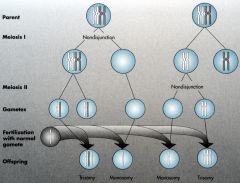
If it occurs during 2nd cell division, two of the four cells will be normal. One will have two chromosomes (receives both chromatids).
|
|
|
Autosome |
A chromosome that isn't sex (X or Y) e.g., a somatic cell |
|
|
Numerical chromosome abnormalities that can exist |
Autosomes: 13, 18, and 31
Sex chromosomes
|
|
|
Aneuploidy |
State of having a chromosome number that's not a multiple of the haploid number.
Loss or gain of a single chromosome.
|
|
|
Euploidy |
Exact multiple of haploid set of chromosomes
Haploid: normal number of gametes (n=23)
Diploid: normal number in zygote and somatic cells (2n = 46)
Polyploidy: complete set(s) of extra chromosomes
|
|
|
Structural Chromosome Abnormalities can be one of two types |
Balanced: no loss or gain of DNA
Unbalanced: abnormalities in DNA
|
|
|
Down syndrome incidence |
As maternal age increases, incidence increases.
Down Syndrome incidence peaks @ 40 years old |
|
|
Chromosome Testing
|
Requirements Must be (1) mitotically active and dividing, and (2) have a nucleus.
Sources
(1st trimester 9-11wks from extraembryonic tissue from chorion)
**Chemicals are used to make mature WBCs divide |
|
|
Amniocentesis |
10-15 mL of amniotic fluid is used.
|
|
|
Procedure for chromosome analysis |
|
|
|
G-Banding Techniques for Chromosomes |
G-Banding: trypsin with Giemsa staining
Different staining pattern (dark and light) based on chromosome contents and structures:
G+ are Dark Bands
G- are Light Bands
|
|
|
Banding for chromosome location: e.g., Band 6p23 |
6p23
Chromosome number = 6 Arm = short (p) Chromosome region = 2 Band in chromosome region = 3
Centromere is considered zero (0) and the number gets higher as you move away from the centromere either up or down the chromosome. |
|
|
Human Karyotyping |
Chromosomes are numbered (1-22) based on their size. They get smaller as the number increases.
Exception: chromosome 22 is larger than 21
Chromosome 21 is smallest chromosome in the human genome. This is why trisomy 21 is most prevalent--because it is most tolerable because it has less genes than other chromosomes. |
|
|
Karyotype Descriptions |
Chromosome #, sex designation, abnormalities
Normal male: 46,XY Normal female: 46,XX
(deletion in short arm of chromosome 5)
- abnormal listed 1st: 47,XX,+21 (trisomy) - normal listed 2nd: 46,XX
|
|
|
Karyotype for the following diseases:
|
Turner: 45,X
Klinefelter: 47,XXY
Down: 47,XY,+21 (if male) |
|
|
Triploidy |
Gain of an extra SET of chromosomes (note this is different than triSOMY)
Types: 69,XXX/XXY/XYY
Symptoms
|
|
|
How many genes are in the human genome? |
22,000 |
|
|
Within each cell, genome is packed as ________. |
Homogenous chromatin (DNA and histones) |
|
|
When cell divides, it's DNA _______ and is visible as chromosome under microscope. |
Condenses |
|
|
Chromosome abnormalities are seen in ____ % of 1st trimester and ____% of 2nd trimester losses. |
1st Trimester: 50%
2nd Trimester: 20% |
|
|
In interphase, chromosome is highly _______ and cannot be visualized. |
Interphase: decondensed (euchromatin)
|
|
|
When nondisjunction occurs during mitosis, one cell gets ____ chromosomes and one cell gets ____ chromosomes. |
One gets 45 and one gets 47.
|
|
|
Remember, Mosaicism is caused by nondisjunction during _____.
It results in ___________. |
MITOSIS
Results in coexistence of two cell populations. |
|
|
Full summary of Meiosis |
Meiosis reduced chromosome number from diploid (46) to hapolid (23). Composed of DNA replication followed by two divisions.
Meiosis I
Meiosis II
|
|
|
Two types of Chromosome Deletions? |
Terminal Deletion: end of telomere is lost.
One breakpoint indicates the terminal segment of chromosome has been lost: 46, XY, del(5)(p14)
Interstitial Deletion: middle of the telomere is lost (two breaks).
e.g., 46,XY,del(5)(p14p15.3) - male with interstitial deletion of 5p segment; only part of short arm of chromosome 5, from p14 o p15.3 is lost.
|
|
|
Both types of chromosome deletions can occur at both mitosis and meiosis.
Unequal crossover will occur ____... |
Unequal crossover occurs at meiosis during pairing and recombination of homologous chromosomes.
|
|
|
Three common deletion syndromes |
1. Wolf-Hirshhorn Syndrome
2. Williams Syndrome
3. Cardiofacial Syndrome
|
|
|
Wolf-Hirshhorn Syndrome 1. Mutation 2. Presentation |
Wolf-Hirshhorn Syndrome
Mutation: terminal del4p
Presentation
|
|
|
Williams Syndrome 1. Mutations 2. Presentation |
Williams Syndrome
Mutation: interstitial del7q11.2
Presentation:
|
|
|
Cardiofacial Syndrome 1. Mutation 2. Presentation |
Mutation: deletion 22q11.2
Most common deletion syndrome in human |
|
|
Isochromosome |
When two sister chromatids break apart at the centromere.
The two short arms combine and the two long arms combine. |
|
|
Ring Chromosome |
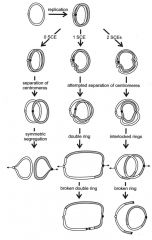
Two terminal ends of chromosome break off and the remaining ends join together to form ring.
Unstable during mitosis. Possibilities:
Ring chromosome phenotype: growth retardation and mild developmental delay |
|
|
Types of Chromosome Inversions (2) |
Both inversions occur without loss or gain of DNA
1. Paracentric: one arm without centromere
2. Pericentric: involves centromere
memory aid:
|
|
|
Paracentric Inversion |
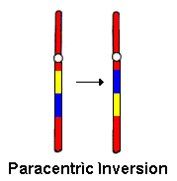
Happens in one arm without involving centromere.
Inversion loop occurs to allow for pairing of homologous chromosomes.
- Dicentric gametes: two centromeres - Acentric gametes: one centromere |
|
|
Pericentric Inversion |
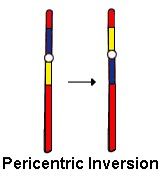
Involves the centromere
Recombination outside inversion loop: normal and inverted
Recombination insider loop leads to four possible gametes:
|
|
|
Difference between peri- and paracentric inversion? |
Products of PERIcentric inversion have ONE centromere and are therefore viable. |
|
|
Reciprocal Translocation |
Breaks at different chromosomes that switch chromosomes.
No loss or gain in genetic material, so they're balanced. Carriers are phenotypically normal but are at risk for having abnormal kids.
One parent contributes two normal chromosomes. Parent with translocation has two options:
|
|
|
Difference between inversions and translocations |
Inversions are at the same chromosome.
Translocations are at different chromosomes. |
|
|
Robertsonian Translocation |
Involves ACROCENTRIC chromosomes (13-15, 21, 22)
Chromosomes join together at the centromere after ends break off of them. This fusion reduces chromosome number from 46 to 45.
This is a balanced translocation and they're phenotypically normal--this is only situation where you can reduce chromosome # from 46 to 45 and be phenotypically normal.
|
|
|
What is the only situation that you can reduce chromosome number from 46 to 45 and still be phenotypically normal? |
Robertsonian translocation involving acrocentric chromosome (13-15, 21, and 22). |
|
|
Translocation-type Down Syndrome (Robertsonian Translocation) |
Carriers of Robertsonian Translocation involving chromosome 21 are at risk for producing child with translocation type down syndrome.
It's important to note that having the Robertsonian Translocation doesn't cause Down Syndrome. It means you're a carrier. It's how this is passed down to the kid that determines whether they'll get it. |
|
|
Female vs. Male Chromosome |
Females down regulate genes on their X-chromosome to equal male levels of expression.
Only one X-chromosome with be active. The other is a Barr body.
Barr Body: inactive X-chromosome in female |
|
|
Lyon hypothesis |
In females, one X chromosome is active.
|
|
|
X-Inactivation is incomplete |
Several genes on inactive X are know to ESCAPE INACTIVATION
|
|
|
Dosage Compensation |
XXY vs XXXY |
|
|
Mosaicism in females |
Each tissue is mosaic in the female.
This is not true mosaicism
Modulational change to control what chromosome is activated or inactivated. It's an EPIGENETIC CHANGE. |
|
|
Variable expression in heterozygote X chromosome |
Diseased X is "turned off" to prevent expression of disease.
Unfortunate Lyonization: mutant allele on active X chromosome (skewed X inactivation)
If there has been a translocation, normal chromosome is inactivated in order to retain the genetic material that has been translocated between the other two chromosomes. |
|
|
Incidence of different Chromosome Abnormalities |
35% = sex chromosome aneuploidies
30% = balanced structural abnormalities
25% = autosomal aneuploidies
10% = unbalanced structural abnormalities |
|
|
What are the names of the three trisomy conditions? |
Trisomy 13: Patau syndrome
Trisomy 18: Edwards syndrome
Trisomy 21: Down syndrome |
|
|
Trisomy 21: Down Syndrome |
Translocation recurrence risk:
|
|
|
Trisomy 18: Edwards Syndrome |
Second most common chromosome abnormality
|
|
|
Trisomy 13: Patau Syndrome |
Third most common chromosome abnormality
|
|
|
Sex Chromosome Anueploidies |
Do better than autosomal aneuploidies because:
Note, some genes on inactive Xs can escape inactivation, so they may not be completely harmless.
|
|
|
Two types of sex aneuploidies? |
1. Turner Syndrome: 45,X
2. Klinefelter Syndrome: 47,XXY |
|
|
Turner Syndrome |
FEMALE ONLY: 45, X
|
|
|
Klinefelter Syndrome |
MALE ONLY: 47,XXY
|
|
|
Genomic Imprinting and two examples |
Genomic Imprinting: certain genes expressed in a parent-of-origin specific manner
1. Prader-Willi syndrome 2. Angleman Syndrome
|
|
|
Prader-Willi Syndrome |
Example of genomic imprinting. Loss of active genes in 15q11-q13 (paternal)
Normally, inherit one copy of chromosome 15 from each parent. Some genes on this chromosome are only active on paternal copy.
|
|
|
Fluorescence in Situ Hybridization (FISH) |
Analyzes chromosome at molecular level
Does not give info about location on chromosome. |
|
|
Conventional Banding Method |
Allows a glance at complete genome in single test.
Limited resolution for detection of small deletions and other subtle rearrangements. |
|
|
DiGeorge Velocardiofacial Syndrom (VCFS) |
Autosomal dominant with VARIABLE EXPRESSIVITY.
Deletion within chromosome 22q11
|
|
|
Preimplantation Genetic Diagnosis with FISH |
Alternative to prenatal diagnosis. Involves selecting preimplantation embryos from cohort generated by assisted reproduction technology.
|
|
|
Array-Based Comparative Genomic Hybridization (aCGH) |
Allows high-resolution scanning of genome.
e.g., label patient DNA green and the control another color. Mix the two and observe whether color mixes appropriately. If preference for one color, microdeletion. |
|
|
FISH vs. Array-Based Comparative Genomic Hybridization (aCGH) |
FISH: one probe
CGH: hundreds of probes
|
|
|
Structural rearrangement should always be analyzed during what cycle of cell? |
Metaphase cells |
|
|
What is gold standard for chromosome analysis and provides view of entire genome? |
G-banding |
|
|
_____ can use a unique-sequence probe to detect microdeletion. |
FISH |
|
|
Multicolor paining allows detection of _____, but not for _______. |
Detection of structural changes.
Not good for small deletions |
|
|
_______-fold improvement between chromosome study and microarray |
1000 fold improvement |

