![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
315 Cards in this Set
- Front
- Back
|
Gram-negative bacteria
|
outer membrane, peptidoglycan layer,
|
|
|
Two main classes of eubacteria
|
gram negative, gram positive
|
|
|
gram-positive bacteria
|
no outer membrane, thicker peptidoglycan layer
|
|
|
What kind of cells have periplasmic space?
|
gram-negative
|
|
|
Explain endosymbiont hypothesis
|
Ancestral eucaryote was anaerobic until it took in an aerobic bacterium. Now eukaryotes can carry out aerobic catabolism. Some bacterial genes moved to the nucleus and the bacteria became mitochondria. For photosynthesis, a photosynthetic bacterium was engulfed and was used to make ATP from sunlight. This bacteria became plastids (chloroplasts).
|
|
|
Four components for a living cell
|
1. Integrity of compartment
2. Energy 3. Respond to environment 4. Reproduce itself |
|
|
Viruses are not alive because ______
|
They do not metabolize.
|
|
|
5 characteristics of eukaryotes
|
1. big
2. have organelles 3. DNA in nucleus 4. Almost always diploid or polyploid 5. Replication via mitosis and meiosis |
|
|
5 characteristics of prokaryotes
|
1. small
2. No organelles 3. DNA is free in cytoplasm 4. Usually haploid 5. Replication via simple division after DNA replication |
|
|
Most common atom in human body
|
H
|
|
|
Most common molecule in E coli
|
Water (70%)
|
|
|
Chiral molecule
|
Rotated molecule can not be superimposed on its mirror image
|
|
|
Enantiomers
|
two mirror image sterioisomers
|
|
|
RS priority system
|
-SH > -OCH2- > -OH > -NH2 > -COOH > -CHO > -CH2OH >
-CH3 > -H |
|
|
Which direction is R?
|
Clockwise
|
|
|
Which direction is S?
|
Counterclockwise
|
|
|
Which direction is L?
|
= S = counterclockwise
|
|
|
Torsion angles have the highest potential energy when
|
the molecule is in eclipsed form and lowest when it's in the staggered formation.
|
|
|
Explain prebiotic evolution
|
Origin of life theories: chemical
evolution from primordial soup simple organic molecules associate into larger complexes spontaneously form membranes and catalysts = precursors to first cells. Miller-Urey (1953) experiments demonstrated feasibility: |
|
|
Describe RNA world hypothesis
|
Prebiotic soup was created, including nucleotides, production of short RNA molecules, selective replication by RNA segments, synthesis of peptides by RNA, coevolution of RNA and protein, genomic RNA copied into DNA.
Evidence is ribosomes and ribozymes. |
|
|
Give a brief definition of a scientific theory; contrast it with an hypothesis.
|
A theory is a model of a portion of nature that accounts for known experimental and
observational facts. A hypothesis is a working guess about nature that can be tested. |
|
|
What are some criterion for experiments to be accepted by the scientific community?
|
Experimental data must be public, controlled, and reproducible.
|
|
|
We wish to make 1 liter of a 0.04 M phosphate buffer at pH = 7.1. What is the ratio of [HPO4-2]/[H2PO4-]? (pK2 (H3
PO4) = 6.8) |
7.1 = 6.8 + log [HPO4^2-]/[H2PO4^-]
0.3 = log [HPO4^2-]/[H2PO4-] [HPO42-]/[H2PO4-] = 2 |
|
|
What concentrations of acid and base are required if [HPO42-] = 2[H2PO4-] from above
|
-[HPO42-] + 2[H2PO4-] = 0
[HPO42-] + [H2PO4-] = 0.04M ______________________ 0 [HPO42-] + 3[H2PO4] = 0.04M 3[H2PO4-] = 0.04M [H2PO4-] = 0.013M [HPO42-] = 0.027M |
|
|
[H2PO4-] = 0.013M
[HPO42-] = 0.027M If a reaction then releases 3 x 10-3 M H+ into the buffer, what is the new pH? |
The acid protonates the base, increasing acid form and decreasing base form.
[H2PO4-] = 0.013 + 0.003 = 0.016 [HPO42-] = 0.027 – 0.003 = 0.024 pH = 6.8 + log 0.024/0.016 = 6.8 + 0.18 = 7.0 |
|
|
The main blood buffer is carbonate. The pK1 for the acid is taken to be 6.1, and
yet the blood is stabilized at pH 7.4. a. Why is the carbonate buffer so effective? |
pH = pKa + log [b]/[a]
7.4 = 6.1 + log [b]/[a] [b]/[a] = 20 The relatively low [acid] is compensated by being generated from CO2 + H2 O Æ HCO3 |
|
|
A patient at a clinic is found to significantly elevated concentrations of H2CO3. Is this
problem respiratory or metabolic? |
Respiratory
|
|
|
A protein has been sequenced and has 250 amino acids. What is the rough
MW? |
110 x 250 = 28,000 Da
|
|
|
What is the significance of the fact that mitochondria have their own DNA?
|
They were once free living bacteria, accreted into higher cells
|
|
|
What about the fact that mitochondria have a double membrane?
|
The double membrane suggests uptake by “endocytosis”
|
|
|
What is the pitch of an α-helix?
|
5.4 angstroms
|
|
|
How many residues are
used to make one turn of the alpha helix? |
3.6
|
|
|
What is the hand of the alpha helix?
|
right
|
|
|
Protein
homologs from different species that have the same function are called |
orthologs
|
|
|
Proteins within the same species that are homologs but
have different functions are called |
paralogs
|
|
|
The similarity between proteins can be judged by aligning their sequences and counting
identical residues; opening “gaps” in the sequences can increase the number of matches, but what must be done to keep the accounting valid? |
In “scoring” an alignment, introducing gaps must be penalized, and more than one or
two novel amino acid matches. |
|
|
Which motif is concatenated in assembling the nucleotide binding domain (also
called the Rossmann fold)? |
βαβ
|
|
|
By what mechanism are protein motifs assemble into physically larger domains?
|
The genes coding for a motif needs to duplicate or fuse with another to express a new,
larger, polypeptide or protein. |
|
|
How does the affinity for O2 compare in the T and R forms?
|
The R-state has a higher affinity for O2 than the T-state
|
|
|
When O2 binds to the first heme iron of deoxy hemoglobin, chemical and structural
changes occur that initiate a shift towards a higher O2 affinity. Briefly describe the two most immediate changes as we discussed in class. |
O2 binding change heme Fe spin state and physical size. It moves into the heme plane,
flattens that plane and pulls the F helix via its covalent link to His F8. |
|
|
What kind of bonds securing the deoxy structure are dramatically broken in the
conversion to the “oxy” state? |
The T state is stabilized by a number of ion pairs (salt bridges) that are broken in the
conversion to the oxy, or R, state. |
|
|
Does BPG stabilize the R or the T state of hemoglobin, and so what?
|
BPG lowers the hemoglobin’s affinity for oxygen by stabilizing the T-state; O2 is more
easily released into the tissue. |
|
|
The Bohr effect describes the binding of O2 to hemeoglobin as a function of pH.
Which quaternary state is stabilized by H+ (by reduced pH). Which state is a stronger acid? |
The Bohr protons stabilize the T state and enhance O2 release (the H+ is bound to His,
but stabilized by the carboxylate in the T-state). The R state, where imidazolium is not so stabilized, is a stronger acid |
|
|
How many antigen binding sites does IgG have?
|
IgG has 2 branches and each branch has a single antigen binding site. Therefore, IgG
has 2 antigen binding sites. |
|
|
Within antibody families, what are hyper variable regions?
|
These are 3 segments of the variable domain that vary radically in amino acid
sequence. They form the antigen binding pocket. |
|
|
What is the peptide chain composition of IgG?
|
L2H2
|
|
|
We developed a simple binding model of O2 to Mb and found the fractional
binding was described by a rectangular hyperbola. Exactly the same formalism works to describe antigen (L) binding to an antibody. That is for this process L + AB ↔ L•AB Θ= (L)/[(L) + 1/Ka] = (L)/[(L) + Kd], where Kd, the dissociation constant is 1/Ka. If we isolate our favorite antibody measure the Kd for a certain antigen ligand as 10-6 M, what (L) will saturate ½ of our antibody? |
L =(Θ/1- Θ)Kd
L =(0.5/1-0.5)*10-6M L = 1x10-6M |
|
|
OK, now a bit harder: what (L) will give 75% saturations? Don’t be afraid to use the
saturation equation if Kd = 10-6 M. |
L =(Θ/1- Θ)Kd
L =(0.75/1-0.75)*10-6M L = 3x10-6M |
|
|
8 kinds of noncovalent interactions
|
charge-charge
charge - dipole dipole - dipole charge-induced dipole dispersion van der waals H bonds |
|
|
Why is water unusual?
|
It has a really melting point and a high boiling point.
|
|
|
What molecules make H bonds? Which pair cant?
|
O - O, N - O
C - O doesn't exist. |
|
|
What does solvation do?
|
Solvation helps protein-ligand interactions. The interaction is stabilized by H-bonding, ionic, and hydrophobic interactions.
|
|
|
isotonic
|
no net water movement
|
|
|
hypertonic
|
water moves out of cells
|
|
|
hypotonic
|
water moves into cells
|
|
|
Define a log
|
Log is the number that 10 has to be raised to to get the answer. So the log of 1000 is 3.
|
|
|
We make a liter of pure water 1 mM in HCl. What is the expected pH of that solution?
|
1 mM = 10^-3 M
pH = -log [10^-3 M] |
|
|
What is Ka?
|
Ka = [products]/[reactants]
|
|
|
How do you find the pKa from the Ka?
|
pKa = -log Ka
|
|
|
How do you find pH from Ka?
|
1. Solve for [H] with the Ka equation:
[H] = Ka[HA]/[A-] 2. Take the negative log of both sides: -log[H] = -log Ka -log[HA]/[A-] 3. Substitute pH and pKa and invert last term: pH = pKa + log[A]/[HA] |
|
|
What is the pH and pKa when 50% of solute dissociates?
|
pH = pKa
|
|
|
What is the Henderson-Hasselbach equation?
|
pH = pKa +log[A-]/[HA]
|
|
|
A weak acid has a pKa =5, we have a solution in which the acid form is 100 mM and base form is 10 mM. What is the pH of the solution?
|
4.
|
|
|
How do you estimate the pKa?
|
pH = 2.87 = -log[H] = 0.13 - 3
[H] = 10^(0.13-3) = 1.34E-3 M = [A] You have 0.1 M of acid 0.1 - 1.34E-3 = 0.0987 M Ka = (1.34E-3)^2/0.0987 pKa = -log Ka = 4.74 |
|
|
What is the buffering region?
|
When 50% is disolved, pH = pKa
|
|
|
When pH is 1 unit above pKa, what is % dissociation?
|
pH = pKa + log[acceptor]/[donor]
pKa + 1 = pKa + log[acceptor]/donor = 10 = 90% dissociated |
|
|
If you want to make 1 liter 0.4 M acetate buffer at pH 5.22 and the pKa for acetic acid is 4.74, how much
|
pH = pKa + log[A]/[HA]
5.22 = 4.72 + log[A]/[HA] log[A]/[HA] = 0.48 10^0.48 = [A]/[HA] = 3 3[HA] = [A] 3[HA] -[A] = 0 1[HA] + [A] = 0.4 = 4[HA] = 0.4/[HA] = 0.1M [A-] = 0.3 M We want 1L 0.1 moles acetic acid and 0.3 moles acetate 0.1 moles = 6.0 g and 2.46 g = acetate |
|
|
How does the blood buffering system work?
|
The carbonate system is the main blood buffering system and has a pKa of 6.1 but a 20x more base than acid. The base comes from CO2 so the CO2 maintains the correct buffer.
|
|
|
conditions affecting [HCO3-] are
|
metabolic
|
|
|
conditions affecting [H2CO3] are
|
respiratory
|
|
|
metabolic acidosis means
|
excess H+, diabetes
|
|
|
metabolic alkalosis means
|
loss of H+, vomiting or poisoning with base
|
|
|
respiratory acidosis means
|
High CO2 which yields [H2CO3], poor clearance of CO2 from lungs
|
|
|
Respiratory alkalosis means
|
rapid removal of CO2, hyperventilation
|
|
|
A patient comes in comatose. Blood gas analyzer shows [HCO3-] = 50 mM and [H2CO3] = 1.4 mM. What's wrong?
|
pH = 6.1 + log 50/1.4 = 7.67 metabolic alkalosis, maybe ingested too much bicarbonate (a Tums addict!)
|
|
|
DNA is ______ charged
|
negatively
|
|
|
Proteins are _______ charged
|
positively
|
|
|
What is the isoelectric point?
|
The pH at which the surface of a molecule carries no charge.
|
|
|
We want to over express our favorite
enzyme, bevoase. Fortunately we have the gene encoding bevoase on a piece of DNA with BamH1 sticky ends. We insert this into the plasmid pBR322 (at right) and transform it into E. coli. How can we identify those transformant colonies that contain the bevoase gene? |
Inserting the bevoase gene at the BamH1 site
Disrupts the tetracycline resistance gene. Therefore, the transformed E. coli will survive when the growth media is supplemented with ampicillin. |
|
|
Consider the following interconversion, which
occurs in glycolysis. Fructose 6-phosphate glucose 6-phosphate; Keq = 1.97 (a) What is G°’ for the reaction (assuming that the temperature is 25 °C)? |
G°’ = -RTlnKeq
= -5.7log(1.97) = -1.7 KJ/mol |
|
|
If the concentration of fructose 6-phosphate is adjusted to 150 mM and that of
glucose 6-phosphate is adjusted to 50 mM, what is change in G? The original change in G is -1.7 KJ/mol. |
G=G°’ + RTlnKeq
= -1.7 KJ/mol + 5.7log(50/150) = -1.7KJ/mol – 2.7 KJ/mol = -4.4 KJ/mol |
|
|
Why are change G°’ and change G different?
|
These values are different because G°’ is based on a fixed concentrations (1M) and
temperature. G is a function of reactant and product concentrations, as well as temperature. |
|
|
Draw the structure of beta-glucose in the pyranose form
|
Look it up.
|
|
|
Draw the C4 epimer of beta-glucose in the pyranose form.
What is the name of this important sugar? |
beta-galactose
|
|
|
Based on what we said in class,
sketch the double reciprocal plot for the inhibition kinetics of the cyclooxygenase enzyme (COX) with ibuprofen. Include at least two lines, one without inhibitor, and one with. |
plot it
|
|
|
What type of inhibitor is ibuprofen?
|
Competitive: ibuprofen binds at the active site of
COX |
|
|
-CH2-CH=CH-CH2- + H2O --> -CH2-CHOH-CH2-CH2- What reaction is this?
|
Lyase
|
|
|
What reaction class is this?
-OOC-CH2-CO-COO- + NADH + H+ ---> -OOC-CH2-CHOH-COO- + NAD+ _ |
oxido-reductase
|
|
|
. Glucose-6-P --> Fructose-6-P
|
Isomerase
|
|
|
Asp-Ser-Phe-Gln + H2O --> Asp-Ser + Phe-Gln
|
Hydrolase
|
|
|
fructose + ATP --> fructose-6-P + ADP
|
Transferase
|
|
|
First law of thermodynamics
|
Molecules want to go from a high to low potential energy state.
|
|
|
What is entropy?
|
S = disorder
Entropy is always increasing in the Universe. S = k ln W Change in S = k ln (Wf/Wi) ; change S = change q/T |
|
|
Enthalpy
|
H = E + PV = the energy of a thermodynamic system
|
|
|
How does free energy relate to enthalpy and entropy?
|
G = H-TS
|
|
|
How does free energy relate to the steady state?
|
G = G^0 + 5.7KJ/mol x logKeq
|
|
|
How do you find the change in free energy of a reaction?
|
change in G = Gp - Gr
|
|
|
How does free energy relate to Keq?
|
change in G = -5.7KJ/mol x log Keq = -1.36 Kcal/mol x log Keq
|
|
|
3 reasons for high energy nature of ATP
|
1. charge separation
2. product stabilization 3. greater hydration |
|
|
coupled reactions
|
using a favorable reaction to drive an unfavorable reaction.
|
|
|
role of NADH
|
NADH, and NADPH, participate in REDOX reactions. These are less familiar than chemical bond making and breaking seen with phosphate transfers.
|
|
|
4 rules for formal oxidation states
|
1.C-C bonds are shared equally
2.Oxygen has FOS = -2; H has FOS =+1 3.More electronegative atoms take formal possession of the shared electrons 4.Sum of formal charges = charge on the molecule |
|
|
oxidation
|
is a loss of electrons, hydrogen, or hydride, or the addition of oxygen.
|
|
|
reduction
|
is a gain of electrons, hydrogen, or hydride, or the loss of oxygen.
|
|
|
The Nernst Equation
|
E’ = E’0 + (RT/nF)ln (ox/red) = E’0 - (RT/nF)ln (red/ox)
Substituting for R, T, F, and converting to base 10 logs, gives: E’ = E’0 + (0.06/n)log (ox/red) |
|
|
How is free energy and the redox potential related?
|
change in G0' = -nFE0'
|
|
|
When change G is negative...
|
E0' is positive.
|
|
|
NADH + H+ --> NAD+ + 2e- +2H+ E0’ = 0.315 V
pyr + 2e- + 2H+ --> lactate E0’ = -0.185 V Find change G. |
NADH + H+ + pyr --> NAD+ + lactate E0’ = 0.130 V
In terms of more conventional free energy measures: G0’ = -nFE0’ = -2 x 96 KJ/molV x 0.13 V = -25 KJ/mol |
|
|
3 components of nucleotide
|
1. nitrogenous base
2. Ribose or deoxyribose sugar 3. Phosphate |
|
|
purines
|
adenine and guanine
|
|
|
pyrimidines
|
cytosine
thymine (DNA) uracil (RNA) |
|
|
2 main sugar ring conformations and where they're found
|
c-2' exo: most common in B DNA
c-3' exo: RNA (A form helix) |
|
|
N-glycosidic bond
|
the bond between the base and the sugar.
It's always in the beta form for natural nucleosides. |
|
|
phospho-diester linkage
|
between the 5' exocyclic carbon and the 3' carbon
|
|
|
Difference between deoxyribose and ribose
|
deoxyribose is missing a hydroxyl on the 2' carbon.
|
|
|
_______ have strong UV absorbance.
|
Nucleic acids
|
|
|
Helix of the B form
|
right-handed
|
|
|
B form DNA
|
right handed
diam: 20 angstroms bp/turn: 10.5 helix rise: 3.4 sugar pucker: c-2' endo |
|
|
A form DNA
|
right handed
diam: 26 A bp/turn: 11 helix rise: 2.6 A sugar pucker: c-3' endo |
|
|
palindrome DNA form
|
inverted repeat on both strands.
|
|
|
mirror repeat DNA form
|
has potential to bind proteins with 2-fold symmetry
|
|
|
Single stranded palindromes can form _______
|
hairpins
|
|
|
structure of tRNA
|
has 2 A-form helices anchored to unusual bond
|
|
|
What do more proteins bind into the major groove?
|
There are four points of reference in the large groove as apposed to three in the small groove. More differentiation.
|
|
|
More GC content =
|
higher melting point
|
|
|
Deamination
|
A genetic mutation. The C is changed to a U and the T binds to it. So a G-C pair because an A-T pair.
|
|
|
Depurination
|
The N-glycosidic bond is hydrolized so the base just pops off.
|
|
|
UV light and mutation
|
The UV light creates two bonds in adjacent thymines and replication can read it. Or it can create another funky bond.
|
|
|
Alkylating agents
|
Nitrosamine is a preservative
|
|
|
Draw these nitrogenous bases
|
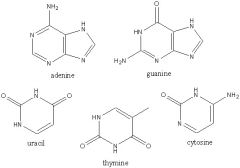
|
|
|
Empirical formula for carbohydrates
|
(CH2O)n
|
|
|
Monosaccharides
|
single polyhydroxy aldehyde or ketone unit (e.g. glucose)
|
|
|
Draw mannose
|
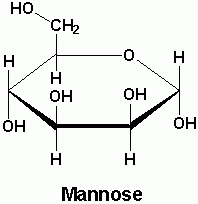
|
|
|
Draw galactose
|
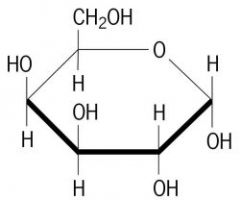
|
|
|
Draw fructose
|

|
|
|
Diasteriomers
|
Stereoisomers with 2 or more chiral centers
|
|
|
anomers
|
a shift of the primary hydroxyl from the beta to alpha position.
stereoisomer at the aromatic carbon |
|
|
Reducing sugar test
|
The ability of the free aldehyde to be oxidized, while something else is reduced, is the basis for most sugar tests.
You react with copper to reduce the sugar by adding O. |
|
|
Epimers
|
Sugars that differ in handedness at ONE non-enatiomeric position
|
|
|
Uronic sugars
|
sugar acids with carbonyl and carboxylic acids
|
|
|
Disaccharides
|
The combination of two monosaccharides that undergo a condensation reaction.
|
|
|
What kind of disaccharide bond can humans digest and the one we cant?
|
alpha bond we can digest
beta bond we can't |
|
|
Draw maltose
|
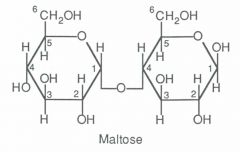
|
|
|
Polysaccharides
|
many monosaccharide units joined in long linear or branched chains. e.g. starch and cellulose (plants) or glycogen (animals)
|
|
|
What is the reducing end?
|
The end of the polysaccharide that can react and be oxidized, ie stuff can be added to it...? Has a potentially free aldehyde.
|
|
|
amylose
|
a component of starch, has a reducing end.
and alpha 1-->4 linkages |
|
|
amylopectin
|
part of starch
linked linearly |
|
|
glycogen
|
animal starch
alpha 1 --> g likages every 10-14 sugars |
|
|
chitin
|
what a bug exoskeleton is made of
beta 1--4 linkages of GlcNAc |
|
|
cellulose
|
largest component of biomass on earth
beta 1 -->4 linkage glucose |
|
|
Heteropolysaccharides
|
2 or more different kinds of monomers
|
|
|
Heteropolysaccharides linked to peptides or proteins. They have differing fractions of carbohydrates and functions.
|
Peptidoglycans
Proteoglycans Glycoproteins |
|
|
Heteropolysaccharides linked to lipids
|
Glycolipids
Lipopolysaccharides |
|
|
Peptidoglycans
|
peptides link carbohydrates
Bacterial cell walls show one example highly glycosylated glycoproteins. They generally have a "core protein" with one or more large glycosaminoglycan (GAG) chain(s) attached, typically at a Ser residue. |
|
|
Proteoglycans
|
often have structural roles: This is a section of bovine cartiledge
|
|
|
Glycoproteins
|
sugars as protein prosthetic groups
o-linked: attached to an oxygen n-linked: attached to N |
|
|
Glycolipids
|
sugars attached to lipids.
Sugars are common cell surface markers. |
|
|
Lectins
|
proteins that bind sugars
|
|
|
zwitterion
|
neutral molecule with a positive and negative charge
|
|
|
isoelectric point
|
pI = (pK1 + pK2)/2
These K's are like the plateau areas. |
|
|
Draw histidine
|
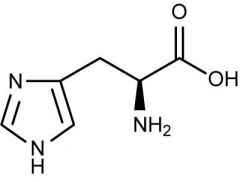
|
|
|
4 ways proteins can be purified
|
1. size
2. charge 3. hydrophobicity 4. binding properties |
|
|
Colormetric assay
|
assay for serine proteases. The fractions that contain the protein turn yellow.
|
|
|
ion exchange chromatography
|
Polymer beads are negatively charged. Large negative proteins move fastest.
|
|
|
Size exclusion (gel filtration) chromatography
|
Porous polymer beads trap small molecules. Big molecules go down first.
|
|
|
Affinity chromatography
|
A mix of proteins are put in the column. Add a ligand and the protein that binds to it will come out first.
|
|
|
SDS-PAGE
|
sodium dodecyl sulfate polyacrylamide gel electrophoresis
The acrylamide gel forms a porous matrix though which proteins can migrate based on charge/mass. In SDS page the proteins are uniformly covered with negatively charged detergent so charge/mass is all the same. The coated proteins move by size through the mesh work; smaller ones run faster. |
|
|
SDS-PAGE with isoelectric point
|
Fill a column with decreasing pI. Then put it in SDS page so that pI decreases to the left and MW decreases down.
|
|
|
Human and bovine cyt c
|
it's 90% identical
|
|
|
What can you use Cyt C for?
|
map evolutionary relationship. The more you have in common, the closer you're related.
|
|
|
orthologs
|
genes in different species that have similar function.
|
|
|
paralogs
|
homologous genes that diverged by gene duplication
|
|
|
The x-ray method
|
1. Grow USEFUL crystals from purified protein
2. Collect X-ray diffraction data; these are only the amplitudes of a complex number 3. The PHASE problem (recover information lost in detection) 4. Build the model, validate it, and interpret the results. |
|
|
Peptide bond
|
Partial double bond prevents rotation of the C-N bond.
Peptide bond is trans. |
|
|
What is the phi bond? The psi bond?
|
phi bond is between the alpha carbon and the N.
Psi bond is between the alpha carbon and another C. |
|
|
How do you calculate the pitch of a an alpha helix?
|
p = n*d
|
|
|
What is the handedness of L amino acids?
|
Right handed
|
|
|
The amino terminus has a ___________ dipole while the carboxyl terminus has a __________ dipole.
|
positive and negative
|
|
|
antiparallel beta sheets
|
hydrogen bonds between O and N are straight
|
|
|
parallel beta sheets
|
H bonds between N and O are sideways
|
|
|
2 types of fibrous proteins
|
1. globular subunits arranged into fibers
2. alpha helical chains twisted together |
|
|
How is alpha keratin made?
|
an alpha helical domain with a globular domain and the alpha domains coil to make a dimer then arrange with other dimers for a protofilament.
|
|
|
Why is hair curly?
|
it gets oxidized - there are fewer disulfide bonds?
|
|
|
collagen
|
right handed triple helical cable - it has a 4-hyp residue. You add a hydroxyl to proline by vitamin c. If you don't have vitamin C, you get scurvy.
|
|
|
4 things on tertiary structure of globular proteins
|
1.Very compact
2.most charged polar R groups (arg, his, lys, asp, glu) are on outer surface, and all are hydrated (i.e. bound water molecules) 3.Most hydrophobic, nonpolar R groups (leu, ile, val, met, phe) buried, hidden from water (blue). 4.Driving force of tertiary structure is partly (mainly) hydrophobic and partly electrostatic. Hydrogen bonding is a strong constraint, but not necessarily a net driving force. Native 3° structure is in dynamic, reversible, equilibrium with other possible conformations, depending of pH, composition, and temp of solution. |
|
|
3 things that stabilize the 3* structure
|
1.Hydrophobic effect
2.Electrostatic forces Dipole-dipole interactions Ionic attractions 3.H-bonding 4.Covalent-linkages |
|
|
Tertiary structure is ultimately determined by
|
aa sequence - denaturation is reversible in many cases. In cases where not reversible, usually large proteins which started folding before completely synthesized - different set of energy minima
|
|
|
Folding rate is much faster than
|
a random process where polypeptide tries all conformations until it finds the right one (1050 years for 100 aa chain). 100 aa synthesized into biologically active protein in 5 sec in E. coli. Must be highly cooperative and ordered.
|
|
|
Domains
|
many polypeptide chains fold into structurally independent globular clusters. Individual domains often have specific functions such as cofactor binding or a particular enzymatic activity
|
|
|
Draw beta-alpha-beta, beta-beta-beta, alpha-alpha, greek key motifs.
|
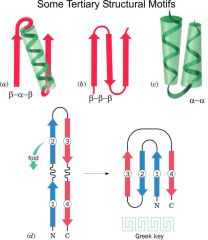
|
|
|
The beta-alpha-beta motif is used in what structure?
|
The Rossman fold which is the NAD binding domain.
|
|
|
Quaternary structure
|
Oligomeric proteins (those with more than one separate polypeptide chain) can range from simple dimer of identical chains to an enzyme complex such as the pyruvate dehydrogenase complex - cluster of 3 enzymes containing 72 chains.
•Protomers are usually symmetrically arranged (cyclic, dihedral, tetrahedral, etc) |
|
|
Name the basic components of the primary, secondary, tertiary, and quaternary structure.
|
primary structure = aa sequence
secondary = conformation of the backbone of the peptide chain tertiary = folding of the secondary structures quaternary = organization of multiple polypeptide chains in a protein complex |
|
|
Most viruses have ______ symmetry
|
icosahedral
|
|
|
Where are the hydrophobic R groups and what are they made of ?
|
They're usually buried in the protein, hiding from water, made up of leu, val, met, phe
|
|
|
_____ is in the center of Heme.
|
Fe^+2
|
|
|
How does heme bind to myoglobin?
|
Heme binds to His F8 in myoglobin. And the O2 is somehow binded to the His E7.
|
|
|
What is the association constant Ka for proteins?
|
Ka = [PL]/[P][L] in M^-1
|
|
|
What is the dissociation constant Kd?
|
Kd = [P][L]/[PL] in M
|
|
|
fractional saturation theta
|
theta = [PL]/[P] + [PL]
Rearrange the Kd: [PL] = [P][L]/Kd theta = [L]/Kd + [L] theta = pO2/P50+pO2 |
|
|
Binding curve for hemoglobin and myoglobin
|
Myoglobin is an exponential curve
Hemoglobin is a sigmoidal curve |
|
|
What does the sigmoidal curve of hemoglobin mean?
|
It means it's cooperative meaning that the affinity changes as more ligands bind.
|
|
|
What is the T state of hemoglobin?
|
It has no oxygen, no binding.
|
|
|
What is the R state of hemoglobin?
|
It is when O is attached.
|
|
|
What stabilizes the T state of hemoglobin?
|
Salt bridges
|
|
|
What is the mechanism for hemoglobin between the T and R state?
|
Binding O causes a conformational change from a dome shape to a flat shape which causes strain on the molecule so His F8 pushes Val FG5 out of the way.
|
|
|
How do you do the hill plot?
|
y axis = log(theta/1-theta)
x axis = log PO2 |
|
|
What other ligands does hemoglobin carry?
|
CO2 and H+
|
|
|
What is the Bohr effect for affinity?
|
pH affects the affinity. For Hb, the higher the pH, the better the affinity.
There's NO effect for Mb. |
|
|
What decreases O2 affinity of the R state in Hb?
|
H+ as well as CO2.
CO2 binding allows new salt bridges to form. |
|
|
What does BPG do?
|
has a higher affinity for deoxygenated Hb. It helps O2 release for RBC.
|
|
|
What's so special about HbF?
|
Fetal Hb has higher affinity for O2.
|
|
|
What causes Sickle Cell anemia?
|
Point mutation where a Glu 6 is changed to a Val.
|
|
|
Describe the structure of IgG
|
4 variable domains.
6 constant domains 1 light chain and 1 heavy chain |
|
|
How many kinds of antibodies are there?
|
4: IgG, IgM, IgD, IgE
|
|
|
What part of the antibody binds the anigen?
|
Fab!
|
|
|
What are IgMs?
|
Membrane-bound proteins that sit on B cells. They have a Y shape with beta sandwiches
|
|
|
Two major components of muscle structure:
|
Actin and myosin
|
|
|
Describe actin
|
Soluble g-actin binds ATP to trigger polymerization. Helix is polar with + at bottom and - at top.
|
|
|
Describe myosin
|
Selective proteolysis can release the globular ATPase heads of the heavy chains; these are called the S1 fragments.
can self assemble into bipolar rods. |
|
|
In a sarcomere, where are the thick and thin filaments and what are the filaments?
|
The filaments are actin and myosin together. The middle is thick only then the really dense part around the middle is thick and thin and then you just have thin.
|
|
|
How do the sarcomeres work during a muscle contraction?
|
The thin and thick filaments come together.
|
|
|
What is the mechanism of contraction?
|
1. ATP binds to myosin head. Actin-binding site releases from myosin.
2. Actin-binding site closes with ATP hydrolysis. 3. Weak binding of myosin head to actin. 4. Pi release makes a strong actin-myosin bond. 5. Muscle contraction makes the unit move. 6. ADP is released. |
|
|
What molecule triggers muscle contraction?
|
Ca2+. It's stored in the sarcoplasmic reticulum and can be released by nerve impulse.
|
|
|
What do microfilaments do?
|
control cell shape and motility.
|
|
|
Microtubules
|
Used in diverse places such as:
omitotic spindle oprotozoan and sperm flagella onerve axons •These systems are constructed from, very long, tubular structures built from a helical arrangement of the protein tubulin. •Composed of two homologous tubulin subunits, a and b, each of molecular weight 55 kDa. |
|
|
Tubulins
|
assemble into microTUBULES
Are tubular, use GTP. Can assemble into the axonemes of cilia or flagella. |
|
|
Dynein
|
In cilia and flagella, used to bend using ATPase activity.
It's in doublets that slide. Moving down the dyenin arms to bend is called "walking". K |
|
|
Bacterial flagella
|
can be made to rotate in either direction and this governs rapid linear movement or tumbling. The cells are observed to move up an attractive gradient by tumbling less frequently and for shorter times.
They rotate circularly. |
|
|
6 facts about Lipids
|
1. energy storage (e.g. triacylglycerols)
2.structural (membranes) 3.Thermal and mechanical insulation 4. Vitamins and hormones (e.g. vitamins A, D; steroids) 5. solubilization of other lipids (e.g. bile acids) 6. insoluable in water |
|
|
Draw palmitic acid
|
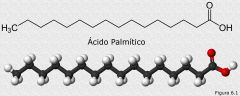
|
|
|
Oleic acid: what's it's melting point?
Draw it. What kind of lipid is it? |
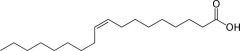
MP = 13.4 C
Unsaturated lipid CH3(CH2)7CH=CH(CH2)7COOH |
|
|
Draw stearic acid. What's the melting point? What kind of lipid is it?
|
Saturated lipid. MW = 69.6 C
CH3(CH2)16COOH |
|
|
In water, lipids spontaneously organize into...
|
micelles and bilayers
|
|
|
What is a wax?
|
An ester between a fatty acid and a fatty alcohol.
|
|
|
triacylglycerols
|
Three fatty acids esterified to the tri alcohol, glycerol.
This is FAT! |
|
|
Fat cells are called...
|
adipocytes
|
|
|
2 kinds of membrane lipids
|
phospholipids
glycolipids |
|
|
2 kinds of phospholipids
|
glycerophospholipids
sphingolipids |
|
|
2 kinds of glycolipids
|
sphingolipids
galactolipids |
|
|
Draw phosphotidic acid.
|
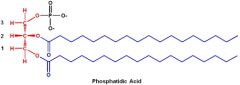
|
|
|
What do sphingolipids have?
|
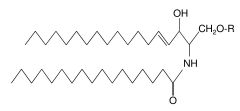
The fatty acid is inked to an amide bond.
|
|
|
What is a ceramide?
|
It is a sphingolipid with an H for the R group.
Diseases like Tay-Sachs arrise from errors in ceramide metabolism. |
|
|
What are sphingolipids used for?
|
blood group antigens A, B, AB, and O.
|
|
|
How do inhibitors work?
|
inhibit COX
|
|
|
Two nonsaponifying lipids
|
steroids
isoprenoids They don't have fatty acids. |
|
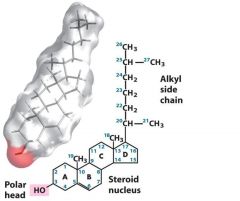
|
Sterol has an alkyl side chain
|
|
|
Steroids
|
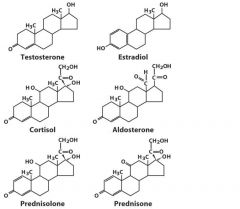
|
|
|
What does it take to be alive?
|
1. Integrity of compartment
2. Energy flow 3. Respond to environment 4. Reproduce itself |
|
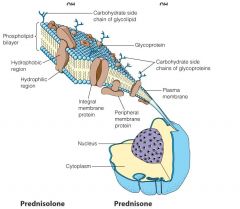
Name all the components
|
Do it.
|
|
|
flippase
|
moves PE and PS from outer to cystolic leaflet
|
|
|
floppase
|
moves phospholipids from cystolic to outer leaflet
|
|
|
scramblase
|
moves lipids in either direction toward equilibrium.
|
|
|
Why is it hard to work with membrane proteins?
|
Peripheral proteins can be taken off with saline, but membrane proteins you have to wash with detergent to coat the protein.
|
|
|
glycophorin
|
a single pass protein with a hydrophobi region.
|
|
|
membrane protein insertion
|
proposed mechanism for a ribosome to find to a cell surface by inserting a peptide into the translocon which moves it.
|
|
|
3 kinds of transport
|
uniport - only one thing goes in
symport - two things go in antiport - one thing goes in and another thing goes out |
|
|
3 mechanisms for facilitated transport
|
1. protein pores
2. carrier molecules 3. permeases |
|
|
ionophores
|
the simplest facilitators
they're carrier molecules |
|
|
GLUT1
|
glucose transporter
permease |
|
|
Equation for Vo
|
Vo = Vmax[S]out/[S]out + Kt
|
|
|
Porins
|
many are helical, but beta barrels are common too.
They are trimers |
|
|
Equation for active transport
|
change G = 5.7(KJ/mol)log(C2/C1) + n96.5(KJ/mol)phi
|
|
|
Na/K ATPase pump
|
It moves three Na out of the cell and 2 K into the cell.
|
|
|
How does the Ach receptor work?
|
Bulky hydrophobic Leu side chain are closed then it binds two Ach inthe M2 helices and then the helices have smaller polar residues and it's open.
|
|
|
DNA sequence with ddNTP
|
Put the mystery sequence in with a bunch of dNTPs and polymerases and ddNTPs. The ddNTPs don't have a 3' OH so it can't attack the alpha-phosphate. You can then take that through a gel to figure out the sequence. READ THE COLUMN FROM BOTTOM TO TOP!
|
|
|
Dye-labeled segments of DNA.
|
Use a primer and add ddNTPs then denature and run through a computer detector to detect what's coming out of the column.
|
|
|
DNA cloning
|
1. Cloning vector cleaved with restriction endonuclease
2. DNA fragments of interest obtained by cleaving chromosome with same endonuclease. 3. Fragments are ligated together. 4. DNA introduced to the host cell. 5. Propogatin (cloning) creates many copies of the recombinant DNA. |
|
|
Type II restriction endonuclease
|
cleaves DNAs at specific base sequences
|
|
|
DNA ligase
|
joins two DNA molecules or fragments
|
|
|
Alkaline phsophatase
|
removes terminal phosphates from either 5' or 3' end
|
|
|
Taq polymerase
|
replicates DNA in PCR reactions
|
|
|
Resistance markers
|
You have to use a resistance marker to make sure your gene of interest has been replicated. So your cell is sensitive to an antibiotic and if you add the vector which as a gene for resistance, the cell becomes resistant.
|
|
|
Lac Z gene
|
The gene metabolizes x-gal which turns blue. If you've correctly inserted that gene, then lac z is interrupted.
|
|
|
DNA amplification
|
1. restriction endonuclease
2. Put foreign DNA fragments into the DNA segment 3. Put it into the phages. |
|
|
Hybridization
|
1. Press nitrocellulose paper into agar plate with bacteria
2. Some cells get on the paper. 3. Treat with alkali to disrupt cells and expose denatured DNA 4. Radiolabeled DNA probe and wash and you can expose x-ray film to paper. |
|
|
Expression vectors
|
Place a gene in a vector by the promotor to sequence that specific protein.
|
|
|
Site-directed mutagenesis
|
You can use a restriction endonuclease or an oligo to put in a new DNA sequence or Add the sequence change with the oligo over replication. It can alter control regions of a gene or change expression.
|
|
|
GST tag
|
You can express a protein tag that's coded along with the DNA sequence and then you run it through a column.
|
|
|
PCR
|
polymerase chain reaction
1. separate ds helix with heat. 2. add oligo 3. Use Taq polymerase to polymerize the two strands 4. Repeat for a long time. |
|
|
Making cDNA
|
You can isolate mRNA and use DNA polymerase I to polymerize the mRNA after a primer to get the cDNA.
|
|
|
Microarray
|
Arrange the DNA and the cDNA on a plate. It will tell you the differences between the two.
|
|
|
oxidoreductases
|
transfer of electrons
|
|
|
transferases
|
group transfer reactions
|
|
|
hydrolases
|
hydrolysis reactions (transfer of functional groups to water)
|
|
|
lyases
|
addition of groups to double bonds, or formation of double bonds by removal of groups
|
|
|
isomerases
|
transfer of groups within molecules to yield isomeric forms.
|
|
|
ligases
|
formation of C-C, C-S, C-O, C-N bonds by condensation reactions coupled with ATP cleavage
|
|
|
ethanol --> acetaldehyde
|
oxidoreductases
|
|
|
Glucose --> G6P
|
Transferases
|
|
|
C-terminus of polypeptide --> shortened polypeptide
|
hydrolases
|
|
|
Pyruvate --> acetaldehyde
|
lyases
|
|
|
Maleate --> fumarate
|
isomerases
|
|
|
pyruvate --> oxaloacetate
|
ligases
|
|
|
What four things do enzymes do?
|
1.Entropy reduction – (proximity and orientation) – enzyme holds substrate(s) so that susceptible bond is in close proximity to catalytic groups.
2.Positions appropriate acid-base groups; some enzymes also use metal ions 3.Desolvation of substrate – Enzyme-substrate replaces most of the water-substrate interactions. 4.Substrate distortion – remember, enzyme binds transition state tighter than ground state. |
|
|
Chymotrypsin
|
hydrolyzed peptide bonds in which carbonyl is donated by aromatic side chain (Trp, Tyr, Phe).
|
|
|
Trypsin
|
is homolog cleaving cationic side chains (Lys, Arg)
|
|
|
Elastase
|
cuts small side chains, like Ala
|
|
|
Subtilisin
|
is bacterial serine protease
|
|
|
Serine proteases
|
are VERY common in living systems for digestion and cell regulation
|
|
|
Serine protease (Cht) active site
|
Ser-195
His-57 Asp-102 |
|
|
Zn++
|
is a cofactor in the mechanism of action of carboxypeptidase
|
|
|
thiamine
|
cofactor: thiamine pyrophosphate
decarboxylation |
|
|
riboflavin
|
flavin mononucleotide
redox |
|
|
niacin
|
nicotinamide
redox |
|
|
pantothenic acid
|
coA
acyl group activation and transfer |
|
|
pyridoxine
|
pyridoxal phosphate
aa activation |
|
|
biotin
|
biotin
CO2 activation and transfer |
|
|
lipoic acid
|
lipoamid
redox |
|
|
folic acid
|
tetrahydrofolate
activation and transfer of C functional groups |
|
|
vitamin B12
|
adenosyl cobalamin
methyl group transfer |
|
|
Draw NAD
|
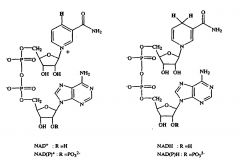
|
|
|
What does Km equal?
|
Km is the Mchaelis constant
Km = k2+k-1/k1 |
|
|
In the double recipricol plot, what is Vmax? What is Km?
|
Vmax is the y intercept.
Km is the x intercept. |
|
|
What is Kcat?
|
The turnover number
kcat = Vmax/[enzyme concentration] |

